TF_ChIP-seq_ENCSR000EVL
| Id: | TF_ChIP-seq/ENCSR000EVL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EVL [biosamplesummary="Homo sapiens HeLa-S3" and target="E2F4"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN981EMX|/analyses/ENCAN981EMX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN981EMX|/analyses/ENCAN981EMX/} has in progress subobject document {d3ffd527-9379-4c6d-894d-404b688a7c68|/documents/d3ffd527-9379-4c6d-894d-404b688a7c68/} audit_not_compliant: Processed alignments file {ENCFF983CXE|/files/ENCFF983CXE/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9566405 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting E2F4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF151MUH|/files/ENCFF151MUH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9014173 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting E2F4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
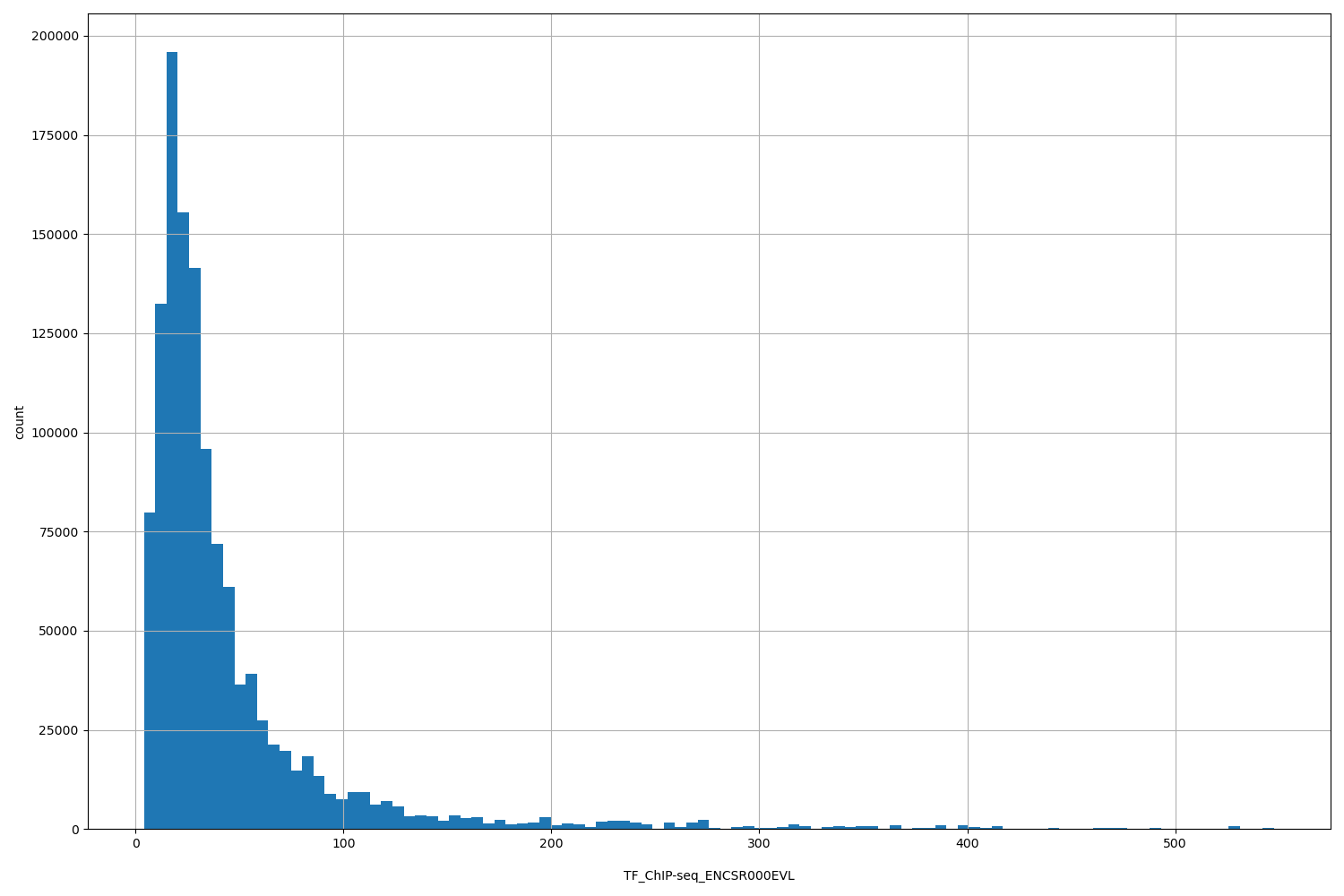
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EVL | float |
TF_ChIP-seq_ENCSR000EVL |
TF_ChIP-seq ENCSR000EVL [biosample_summary="Homo sapiens HeLa-S3" and target="E2F4"]
|
 |
[4, 548] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF718JTU.bed.gz | 59.45 KB | ad352fe9e0e78f8630403b52e2c274fc |
| ENCFF718JTU.bed.gz.dvc | 99.0 B | 1c54c72790a0efced63e40f867445dfb |
| ENCFF718JTU.tabix.bed.gz | 44.82 KB | b01c3e433fb51793e74a308a9489086f |
| ENCFF718JTU.tabix.bed.gz.dvc | 105.0 B | 506fbd762eb71535f30c2aee3bd72937 |
| ENCFF718JTU.tabix.bed.gz.tbi | 32.23 KB | 2f6249cfce95e4ca09e223fd91f47b2b |
| ENCFF718JTU.tabix.bed.gz.tbi.dvc | 109.0 B | d3979454cf8338649f2c2fcd1974f4b1 |
| genomic_resource.yaml | 2.58 KB | 382ed90072c6ba308295412a0551f645 |
| genomic_resource_original.yaml | 2.48 KB | 296806dd56b383c3276d6922b7342e21 |
| statistics/ |