TF_ChIP-seq_ENCSR000EUP
| Id: | TF_ChIP-seq/ENCSR000EUP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EUP [biosamplesummary="Homo sapiens H1" and target="MAX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_error: Processed alignments file {ENCFF764OCO|/files/ENCFF764OCO/} processed by ChIP-seq ENCODE3 hg19 pipeline has 4271427 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF481XFG|/files/ENCFF481XFG/} processed by ChIP-seq ENCODE3 hg19 pipeline has 3699012 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Archived analysis {ENCAN437MBR|/analyses/ENCAN437MBR/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF548MEP|/files/ENCFF548MEP/}, {ENCFF668VNS|/files/ENCFF668VNS/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.16 and a self consistency ratio of 2.86. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF657ICR|/files/ENCFF657ICR/}, {ENCFF268FDZ|/files/ENCFF268FDZ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.16 and a self consistency ratio of 2.86. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
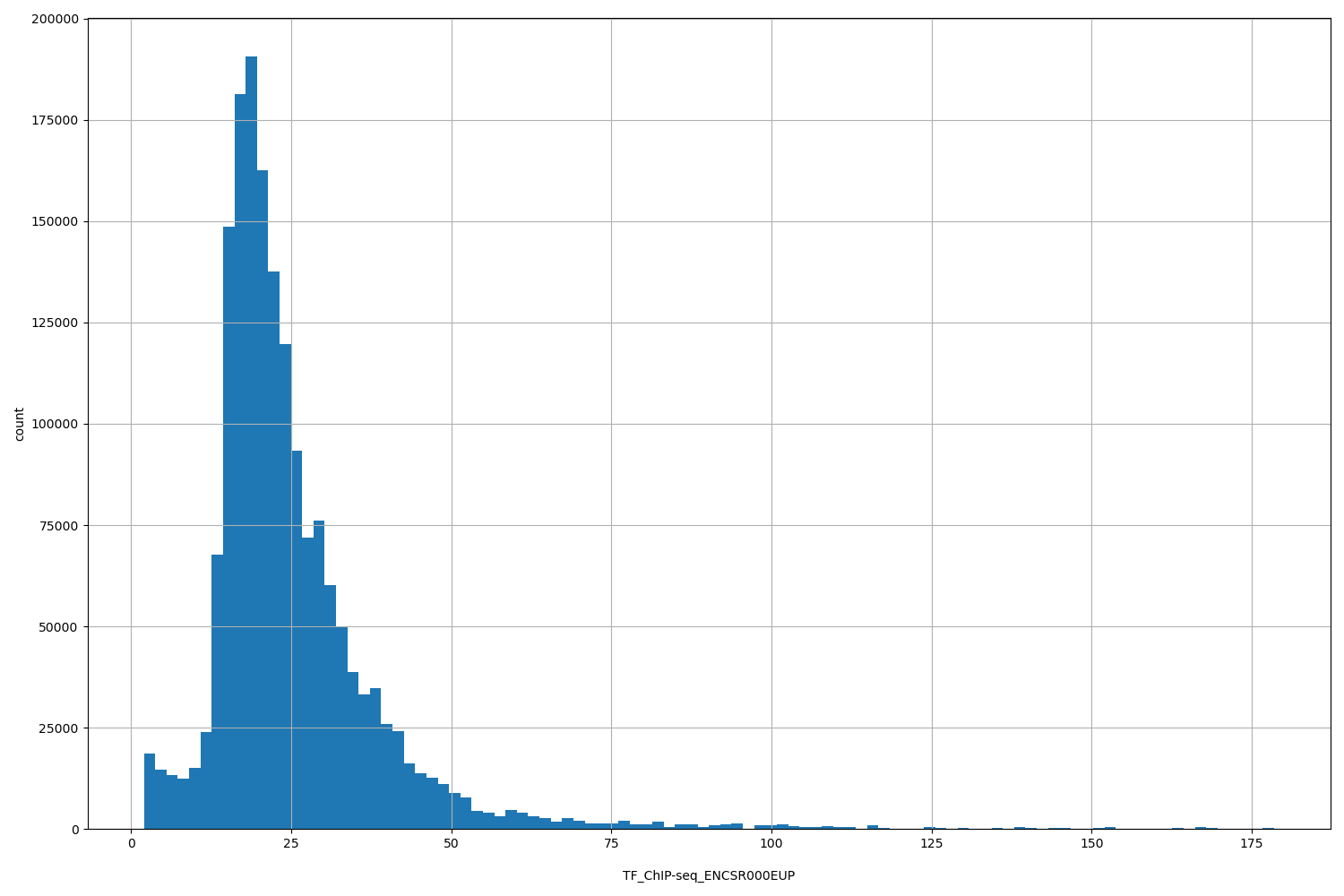
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EUP | float |
TF_ChIP-seq_ENCSR000EUP |
TF_ChIP-seq ENCSR000EUP [biosample_summary="Homo sapiens H1" and target="MAX"]
|
 |
[2.01, 178] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF668VNS.bed.gz | 92.35 KB | d5e88944c656337613f322a78d3b9172 |
| ENCFF668VNS.bed.gz.dvc | 99.0 B | c6701c9de16888201580f3920267f291 |
| ENCFF668VNS.tabix.bed.gz | 71.14 KB | f3bf1e82a0aff7d20bd315d80f5296e2 |
| ENCFF668VNS.tabix.bed.gz.dvc | 105.0 B | c8fe164cd92561638d1a53e248bd87e8 |
| ENCFF668VNS.tabix.bed.gz.tbi | 55.57 KB | 1b9f176eb806dfa87163ecbbd1cced2a |
| ENCFF668VNS.tabix.bed.gz.tbi.dvc | 109.0 B | b3f3cba032ad5218d3ad01c02bf5039b |
| genomic_resource.yaml | 3.44 KB | efbe25ea13d263d19b8757462b70b565 |
| genomic_resource_original.yaml | 3.35 KB | a3f9d8073c932fe6b84284918dedadfd |
| statistics/ |