TF_ChIP-seq_ENCSR000EUN
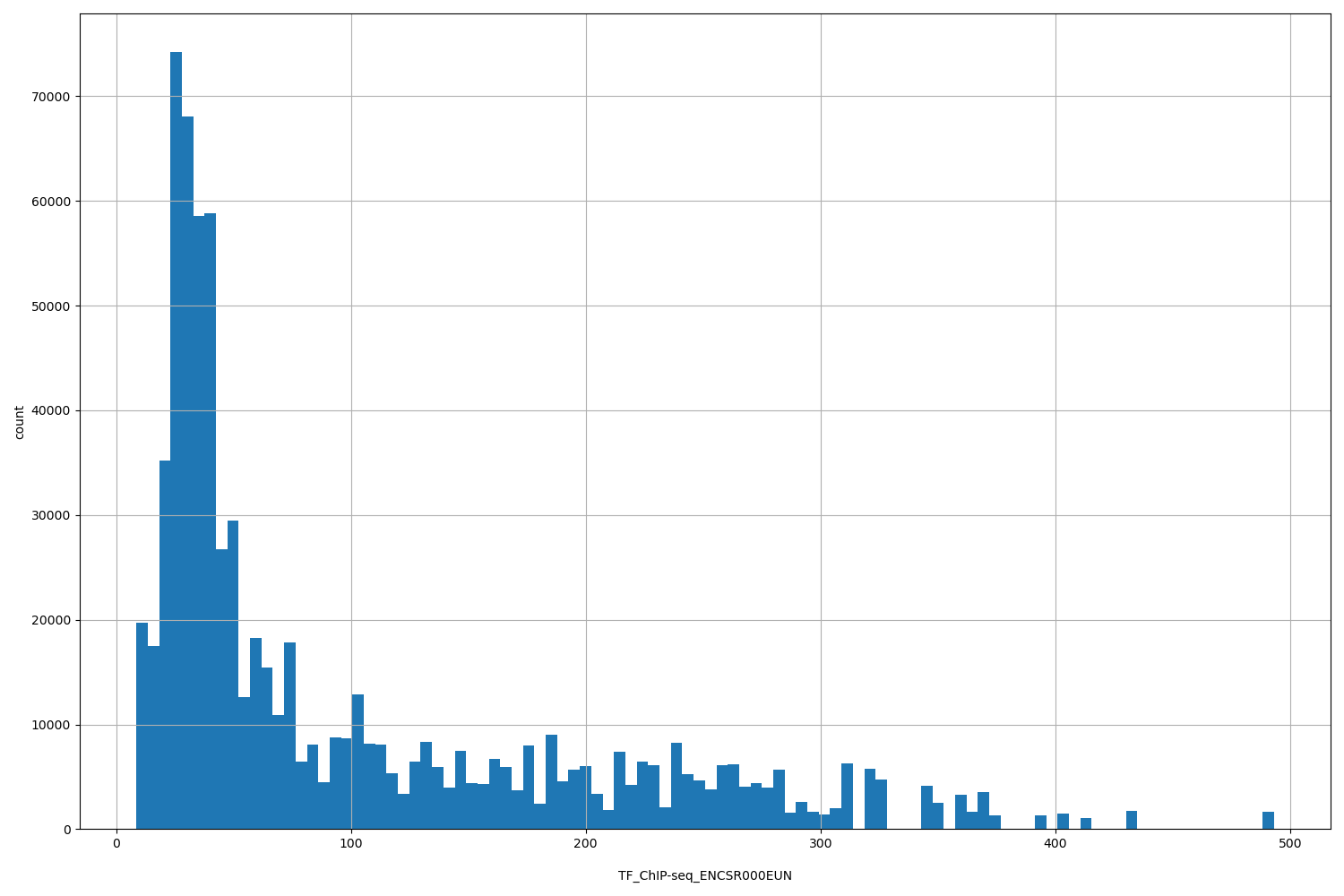
| Id: | TF_ChIP-seq/ENCSR000EUN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EUN [biosamplesummary="Homo sapiens H1" and target="ZNF274"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF943NOT|/files/ENCFF943NOT/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN649QCL|/analyses/ENCAN649QCL/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_error: Processed alignments file {ENCFF101DVO|/files/ENCFF101DVO/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 2964988 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF274-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF943NOT|/files/ENCFF943NOT/}, {ENCFF239NXT|/files/ENCFF239NXT/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.85. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF192YEK|/files/ENCFF192YEK/}, {ENCFF655EHG|/files/ENCFF655EHG/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.85. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EUN | float |
TF_ChIP-seq_ENCSR000EUN |
TF_ChIP-seq ENCSR000EUN [biosample_summary="Homo sapiens H1" and target="ZNF274"]
|
 |
[8.55, 493] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF943NOT.bed.gz | 24.53 KB | 9aeebb6c94faeda4fc0a3408ec829fa1 |
| ENCFF943NOT.bed.gz.dvc | 99.0 B | 5747489575c264c6e22d19a06c0c5fab |
| ENCFF943NOT.tabix.bed.gz | 15.62 KB | da39f5e57cb70ed9a70e5c13d256e296 |
| ENCFF943NOT.tabix.bed.gz.dvc | 105.0 B | 94523a00e09e19dcb119cb6fba7f1f7d |
| ENCFF943NOT.tabix.bed.gz.tbi | 13.22 KB | 2558cb6d2490311253e3dd7f9de87713 |
| ENCFF943NOT.tabix.bed.gz.tbi.dvc | 109.0 B | 93f254c6504e8f740f010e01be8886e1 |
| genomic_resource.yaml | 3.15 KB | 3421e3e35fca6f6a7bc37098bae2db2c |
| genomic_resource_original.yaml | 3.06 KB | fb938591bbe87fe52af078b5c68aa59b |
| statistics/ |