TF_ChIP-seq_ENCSR000EUL
| Id: | TF_ChIP-seq/ENCSR000EUL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EUL [biosamplesummary="Homo sapiens GM12878" and target="NR2C2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF434HVY|/files/ENCFF434HVY/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN095EHD|/analyses/ENCAN095EHD/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF199XSZ|/files/ENCFF199XSZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 12310805 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NR2C2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF485BYB|/files/ENCFF485BYB/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 14252292 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NR2C2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF434HVY|/files/ENCFF434HVY/}, {ENCFF892GAT|/files/ENCFF892GAT/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.43 and a self consistency ratio of 2.27. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF261WWV|/files/ENCFF261WWV/}, {ENCFF992PRK|/files/ENCFF992PRK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.43 and a self consistency ratio of 2.27. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
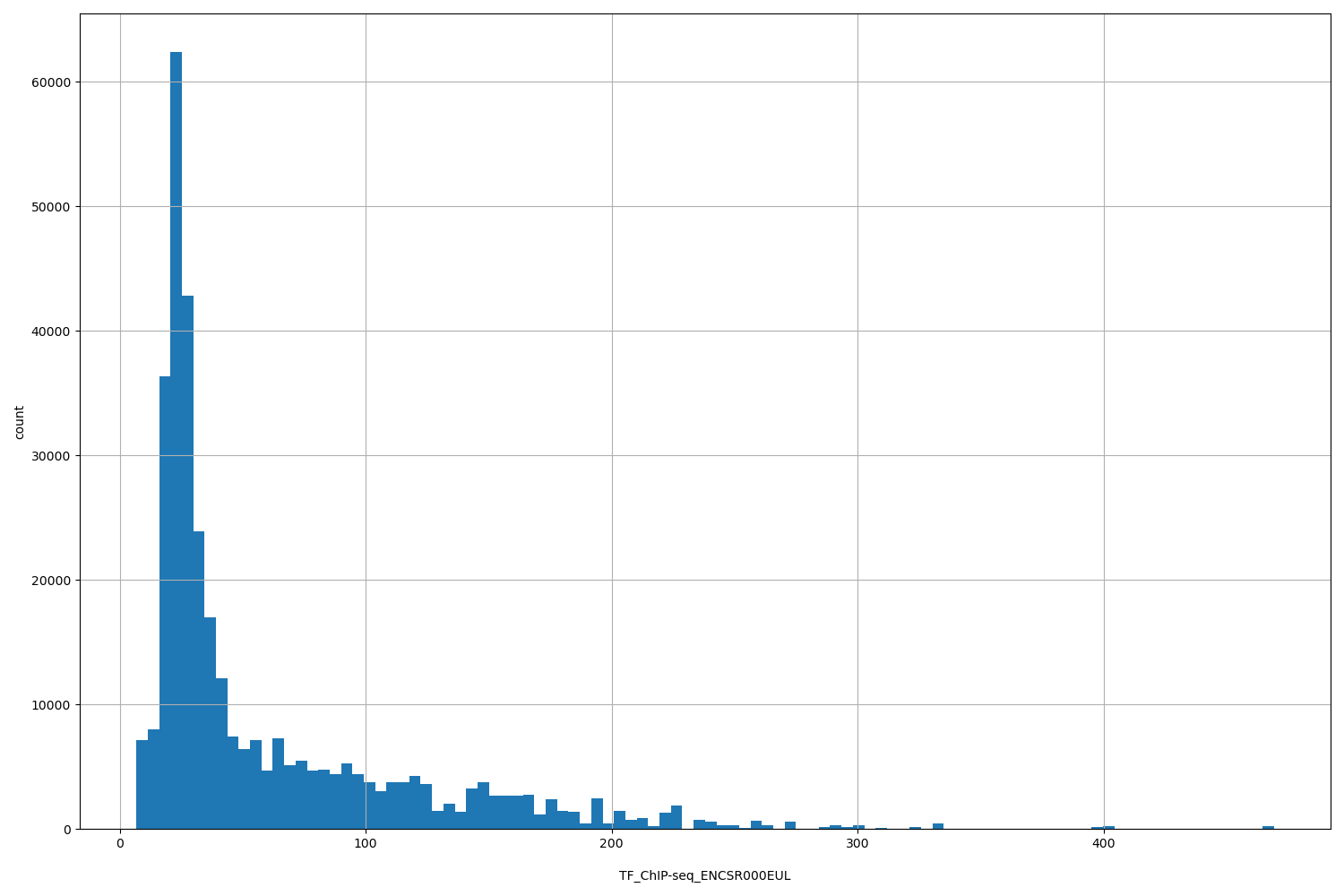
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EUL | float |
TF_ChIP-seq_ENCSR000EUL |
TF_ChIP-seq ENCSR000EUL [biosample_summary="Homo sapiens GM12878" and target="NR2C2"]
|
 |
[6.77, 469] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF434HVY.bed.gz | 19.78 KB | b777a21430fdf491cb5da540cc6afe62 |
| ENCFF434HVY.bed.gz.dvc | 99.0 B | 262b52a3fae9a63a3d9f83f21a73db27 |
| ENCFF434HVY.tabix.bed.gz | 13.66 KB | fc19bfe59cfa193ed408338ec81ecbd9 |
| ENCFF434HVY.tabix.bed.gz.dvc | 105.0 B | 48c3af0b45ddd33c62fc22fdc5c28990 |
| ENCFF434HVY.tabix.bed.gz.tbi | 15.51 KB | 184df6bfb2daea979e5a17348f1daf6e |
| ENCFF434HVY.tabix.bed.gz.tbi.dvc | 109.0 B | fa2f99d5fdbe051bfc11bf301695c5df |
| genomic_resource.yaml | 3.66 KB | ffb06ac3e53ba417b059ac34ed32d637 |
| genomic_resource_original.yaml | 3.56 KB | c65d4f7edb9ac5993efe214414e70555 |
| statistics/ |