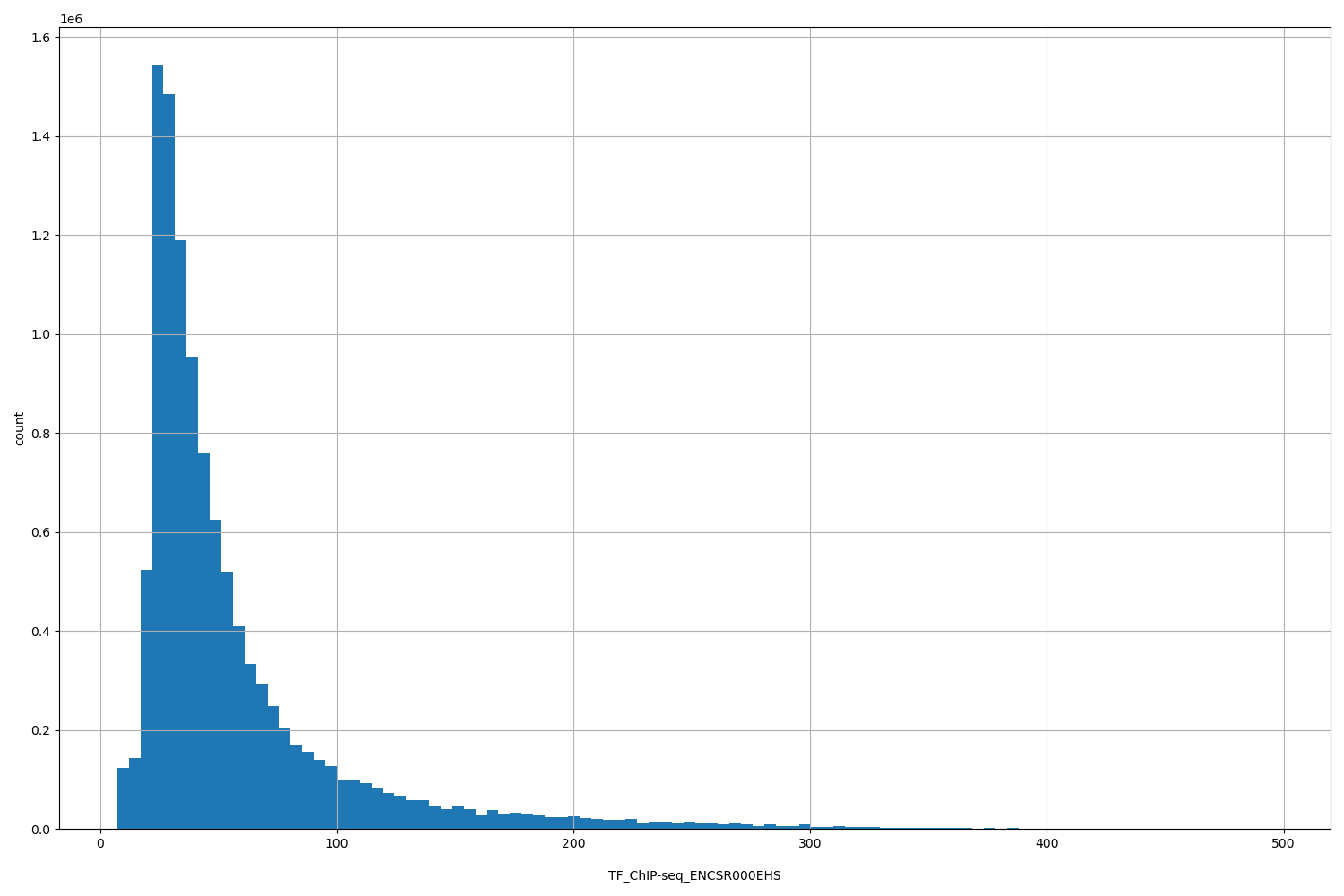
TF_ChIP-seq_ENCSR000EHS
| Id: | TF_ChIP-seq/ENCSR000EHS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EHS [biosamplesummary="Homo sapiens NB4" and target="MAX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN350UOX|/analyses/ENCAN350UOX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN350UOX|/analyses/ENCAN350UOX/} has in progress subobject document {67e8fe39-b1de-428d-8390-f8e8c121657a|/documents/67e8fe39-b1de-428d-8390-f8e8c121657a/} audit_warning: Processed alignments file {ENCFF699XVF|/files/ENCFF699XVF/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19718576 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF044HPR|/files/ENCFF044HPR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19551211 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EHS | float |
TF_ChIP-seq_ENCSR000EHS |
TF_ChIP-seq ENCSR000EHS [biosample_summary="Homo sapiens NB4" and target="MAX"]
|
 |
[7.14, 495] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF793GVV.bed.gz | 691.59 KB | c8fb46fe18bd4a3a65db4ff0f556774e |
| ENCFF793GVV.bed.gz.dvc | 100.0 B | 2f300b2fac5f7f964537656447eb0b45 |
| ENCFF793GVV.tabix.bed.gz | 492.03 KB | 30ada28ccacd7f376a5453fac54ee22a |
| ENCFF793GVV.tabix.bed.gz.dvc | 106.0 B | cc94daaede9d96e55cf208b0c1e4d843 |
| ENCFF793GVV.tabix.bed.gz.tbi | 220.79 KB | 088d9c79504cd4249e1098416682035d |
| ENCFF793GVV.tabix.bed.gz.tbi.dvc | 110.0 B | 16b63a64ef56a8e70b3f9743607198c1 |
| genomic_resource.yaml | 2.55 KB | 787a1d0c2873cef313daf47121078775 |
| genomic_resource_original.yaml | 2.46 KB | 71623ef14bb8615cd7e117312e499957 |
| statistics/ |