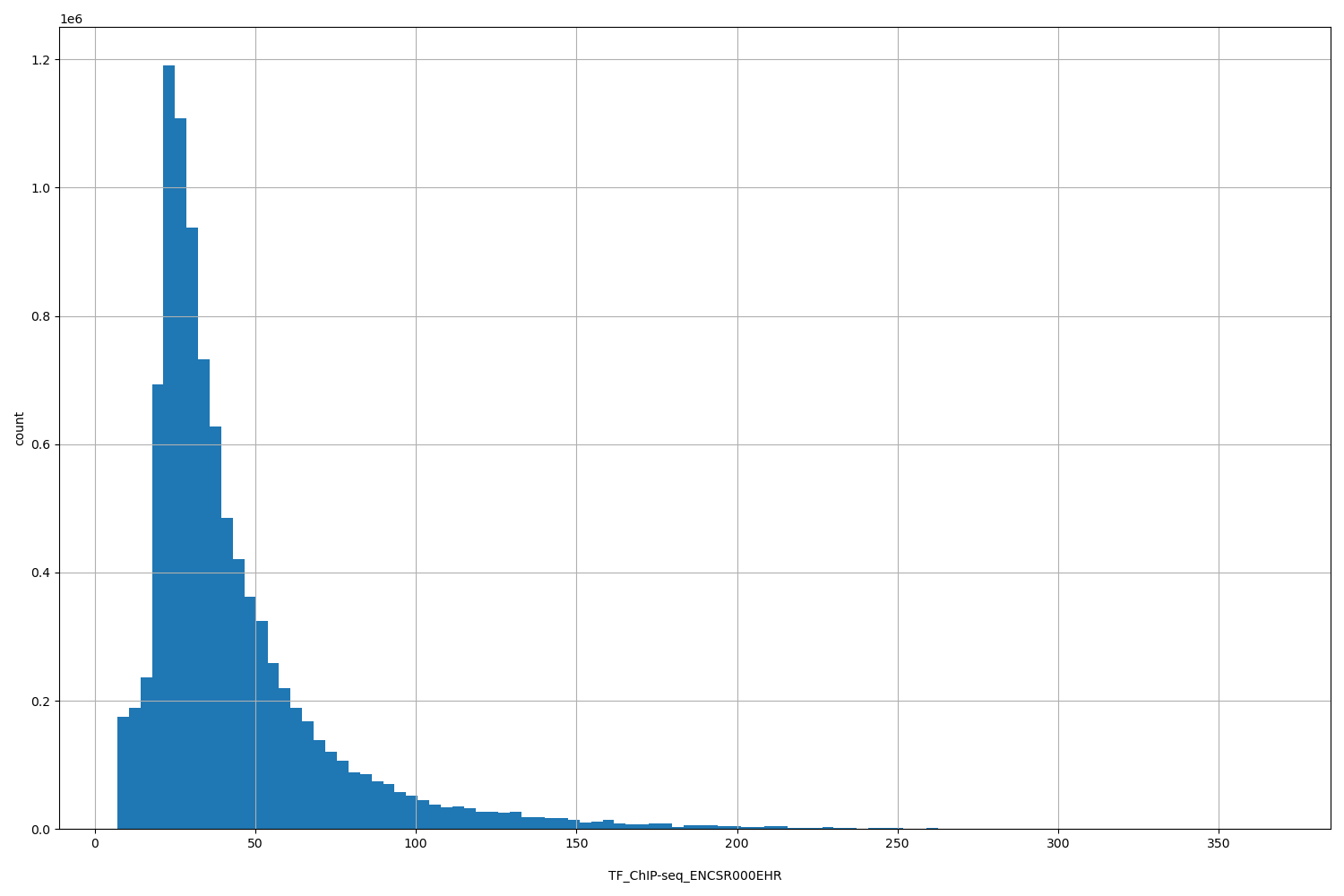
TF_ChIP-seq_ENCSR000EHR
| Id: | TF_ChIP-seq/ENCSR000EHR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EHR [biosamplesummary="Homo sapiens NB4" and target="MYC"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN469SKT|/analyses/ENCAN469SKT/} has in progress subobject document {f834cd5d-b806-4db7-8f1c-2dc4fd1cf59e|/documents/f834cd5d-b806-4db7-8f1c-2dc4fd1cf59e/} audit_internal_action: Released analysis {ENCAN469SKT|/analyses/ENCAN469SKT/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF179WPB|/files/ENCFF179WPB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18818191 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF162XEY|/files/ENCFF162XEY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18243096 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EHR | float |
TF_ChIP-seq_ENCSR000EHR |
TF_ChIP-seq ENCSR000EHR [biosample_summary="Homo sapiens NB4" and target="MYC"]
|
 |
[7, 367] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF858ZYN.bed.gz | 535.54 KB | c30eb368ca462335889995e2397e1786 |
| ENCFF858ZYN.bed.gz.dvc | 100.0 B | dfbf3bf88aae29f638e3e013b95fd454 |
| ENCFF858ZYN.tabix.bed.gz | 383.76 KB | 7df924f6ced2b814fa3ccd37f41f4946 |
| ENCFF858ZYN.tabix.bed.gz.dvc | 106.0 B | faed43649f292419fc041343dec78c57 |
| ENCFF858ZYN.tabix.bed.gz.tbi | 169.52 KB | 694fed9711ef0e80af51b0ce16712e38 |
| ENCFF858ZYN.tabix.bed.gz.tbi.dvc | 110.0 B | 23944fc3dcd78fe187abf38f5f856edd |
| genomic_resource.yaml | 2.55 KB | 143de9da60aa8eecfe611a3046908030 |
| genomic_resource_original.yaml | 2.46 KB | 2e5025d6da15d624a68f196831523f3f |
| statistics/ |