TF_ChIP-seq_ENCSR000EHG
| Id: | TF_ChIP-seq/ENCSR000EHG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EHG [biosamplesummary="Homo sapiens K562" and target="USF2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN761UJX|/analyses/ENCAN761UJX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN761UJX|/analyses/ENCAN761UJX/} has in progress subobject document {f08c84da-9291-4b75-80b1-0b1041991f8b|/documents/f08c84da-9291-4b75-80b1-0b1041991f8b/} audit_warning: Processed alignments file {ENCFF526PJI|/files/ENCFF526PJI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16359580 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting USF2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF332TRZ|/files/ENCFF332TRZ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16163762 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting USF2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
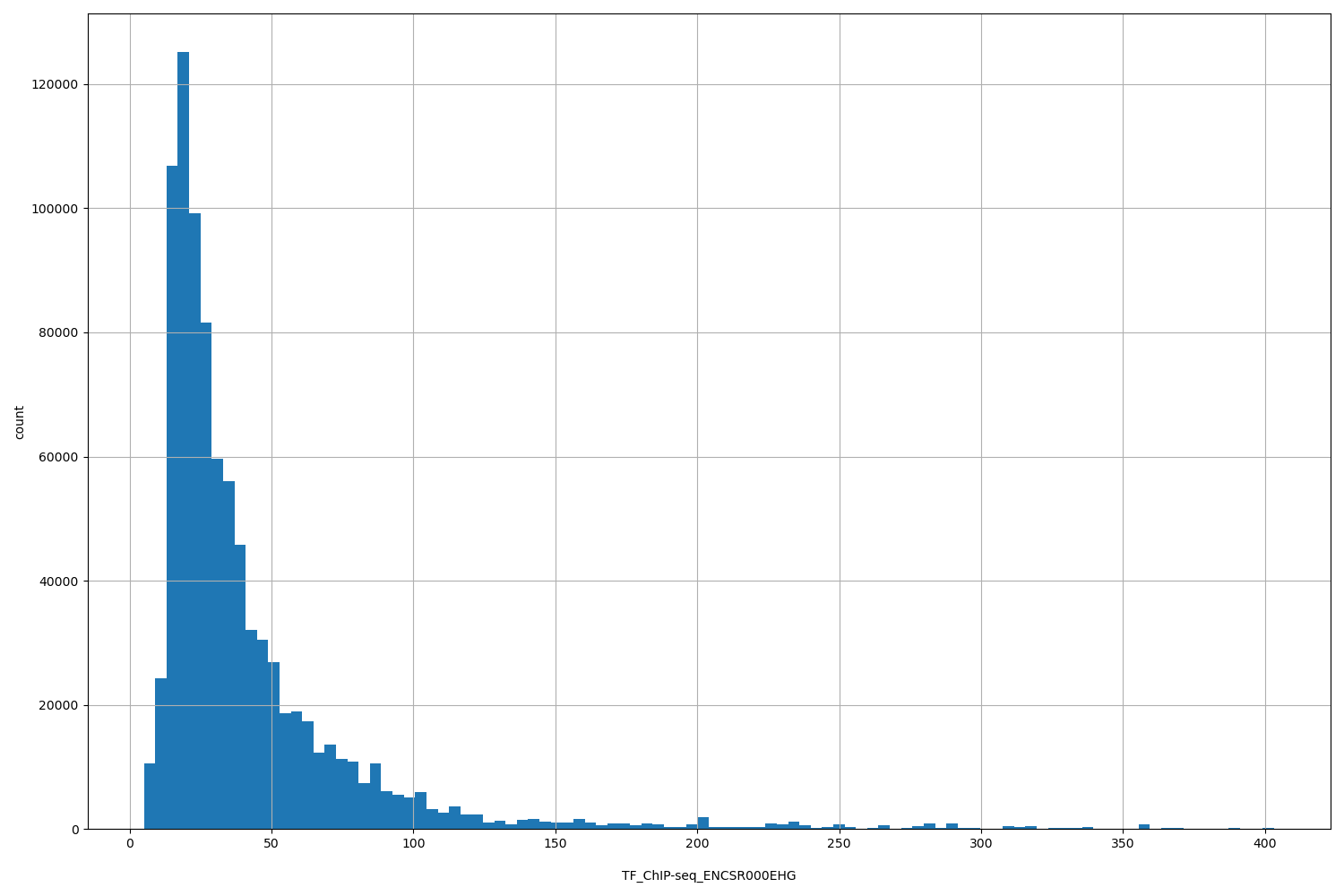
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EHG | float |
TF_ChIP-seq_ENCSR000EHG |
TF_ChIP-seq ENCSR000EHG [biosample_summary="Homo sapiens K562" and target="USF2"]
|
 |
[5.03, 403] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF744HVD.bed.gz | 66.14 KB | 9e1e6d57dfac1bad9fa19e5a3c14b955 |
| ENCFF744HVD.bed.gz.dvc | 99.0 B | a7911ad544c111fabb0dd9abd5007314 |
| ENCFF744HVD.tabix.bed.gz | 43.66 KB | ef449dbe749e0549d75a6d4337b52928 |
| ENCFF744HVD.tabix.bed.gz.dvc | 105.0 B | e791b63c15feafa5ddbf912eafe2772f |
| ENCFF744HVD.tabix.bed.gz.tbi | 37.21 KB | 47b5793bb709fdd5a5f5b405a136f4b3 |
| ENCFF744HVD.tabix.bed.gz.tbi.dvc | 109.0 B | dbb39fa8214a96dbbb3628850ed2bbb9 |
| genomic_resource.yaml | 2.55 KB | 48c02fccb8d72d9020df6d71fb366103 |
| genomic_resource_original.yaml | 2.46 KB | a52b5fa59b8eea6c4ee1b341f3040699 |
| statistics/ |