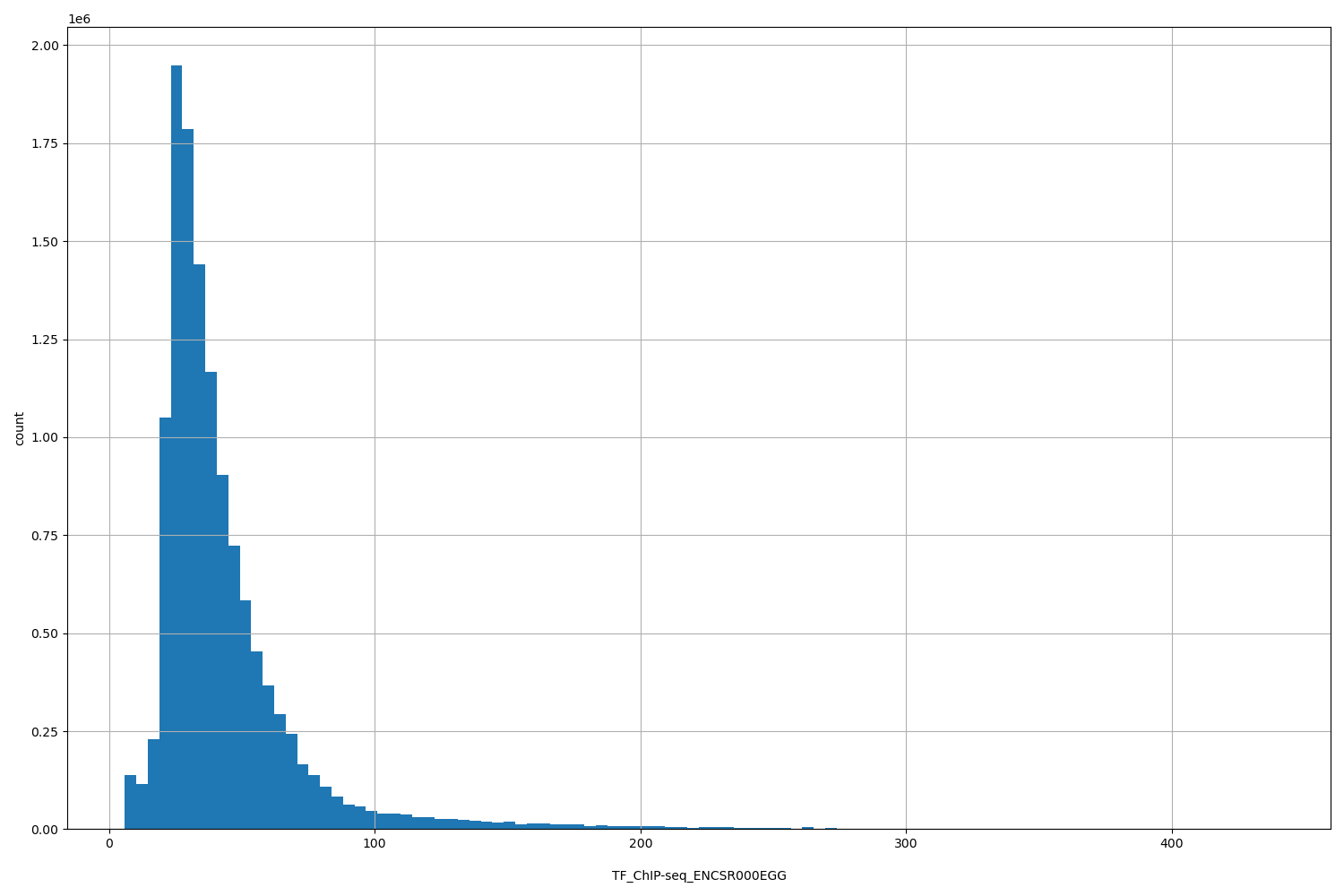
TF_ChIP-seq_ENCSR000EGG
| Id: | TF_ChIP-seq/ENCSR000EGG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EGG [biosamplesummary="Homo sapiens K562" and target="RCOR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN130TEN|/analyses/ENCAN130TEN/} has in progress subobject document {f93d37b2-1f67-401a-842a-d2ae9b148294|/documents/f93d37b2-1f67-401a-842a-d2ae9b148294/} audit_internal_action: Released analysis {ENCAN130TEN|/analyses/ENCAN130TEN/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF557SPX|/files/ENCFF557SPX/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19386085 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting RCOR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF619EXP|/files/ENCFF619EXP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16390218 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting RCOR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EGG | float |
TF_ChIP-seq_ENCSR000EGG |
TF_ChIP-seq ENCSR000EGG [biosample_summary="Homo sapiens K562" and target="RCOR1"]
|
 |
[5.99, 438] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF627PLM.bed.gz | 727.48 KB | 4f39e90d8f3d3a7f7c7f112c8c00cec8 |
| ENCFF627PLM.bed.gz.dvc | 100.0 B | 0f170452a2444eb817d88ae8b7981428 |
| ENCFF627PLM.tabix.bed.gz | 533.08 KB | 5526fc17d0ffe6b8947193a8713c6cf9 |
| ENCFF627PLM.tabix.bed.gz.dvc | 106.0 B | e5ea617f2b5cebc187be10451d9559d5 |
| ENCFF627PLM.tabix.bed.gz.tbi | 224.53 KB | dc438af83b77f4be184c2a263bc89059 |
| ENCFF627PLM.tabix.bed.gz.tbi.dvc | 110.0 B | 6728bff64ef4751382ce69ab23009b75 |
| genomic_resource.yaml | 2.56 KB | 341d2a21d7decbcccfa41f06253b60ad |
| genomic_resource_original.yaml | 2.46 KB | b8eac57f3ef99d89ca636b6d58601adc |
| statistics/ |