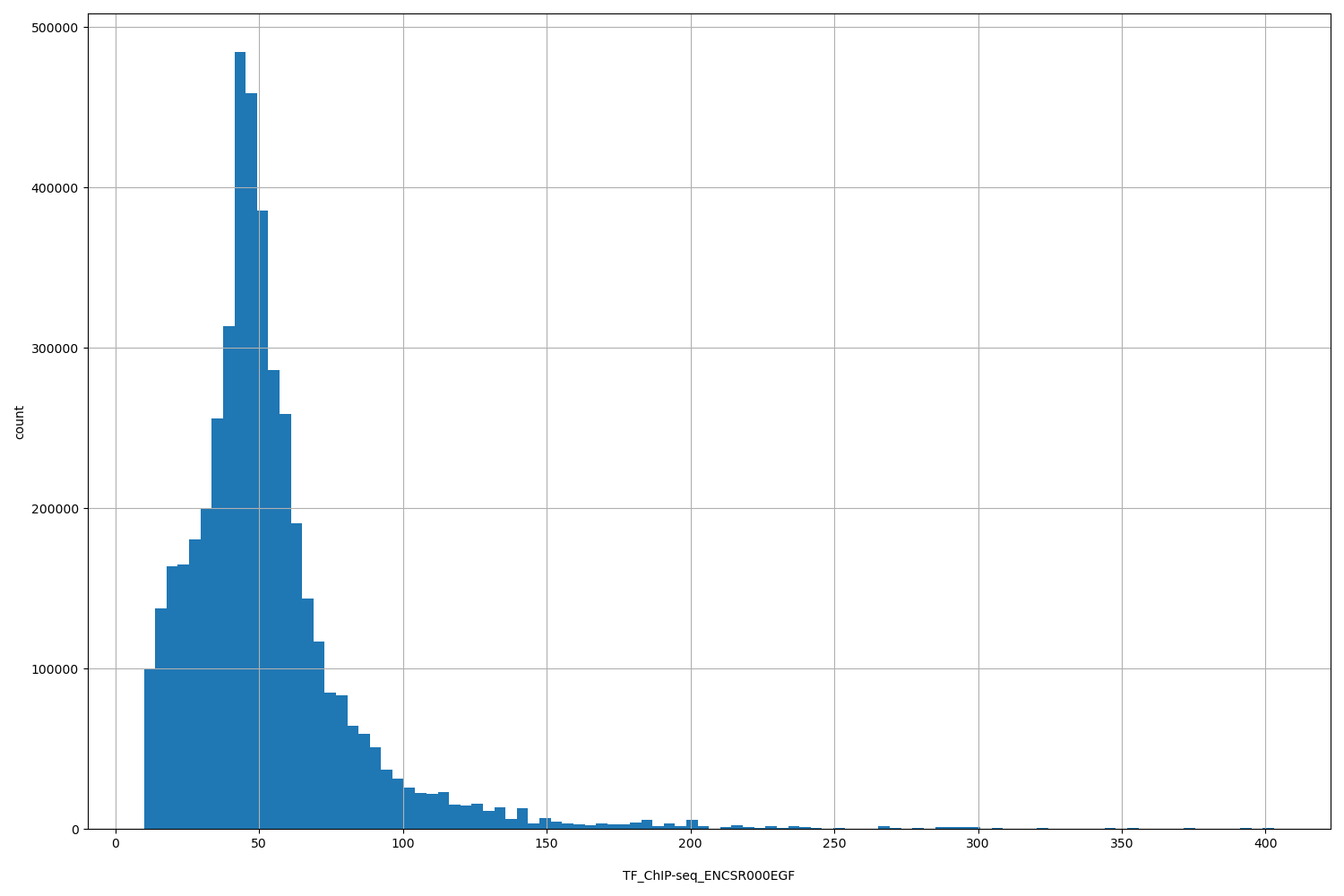
TF_ChIP-seq_ENCSR000EGF
| Id: | TF_ChIP-seq/ENCSR000EGF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EGF [biosamplesummary="Homo sapiens K562" and target="POLR2AphosphoS2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN584UAK|/analyses/ENCAN584UAK/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF207UON|/files/ENCFF207UON/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19700415 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2AphosphoS2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EGF | float |
TF_ChIP-seq_ENCSR000EGF |
TF_ChIP-seq ENCSR000EGF [biosample_summary="Homo sapiens K562" and target="POLR2AphosphoS2"]
|
 |
[9.89, 403] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF296CNQ.bed.gz | 181.34 KB | 96912e3d9f88029951abcd019f3a5721 |
| ENCFF296CNQ.bed.gz.dvc | 100.0 B | a5d8b00142d60ef3338a1f943f776dbe |
| ENCFF296CNQ.tabix.bed.gz | 119.73 KB | bff4a5df8a0465a398c482115502e463 |
| ENCFF296CNQ.tabix.bed.gz.dvc | 106.0 B | 75598434b0c1c1ec311a6b78c94d6b69 |
| ENCFF296CNQ.tabix.bed.gz.tbi | 51.29 KB | bd0ac923d287a662d889e653c565db88 |
| ENCFF296CNQ.tabix.bed.gz.tbi.dvc | 109.0 B | 773778cceab6aa1047daf6cd06b29023 |
| genomic_resource.yaml | 1.88 KB | 36c203e05f36fd0c2543ed20a0e63bc1 |
| genomic_resource_original.yaml | 1.78 KB | 2a860946898e4db68dfba10632596b0f |
| statistics/ |