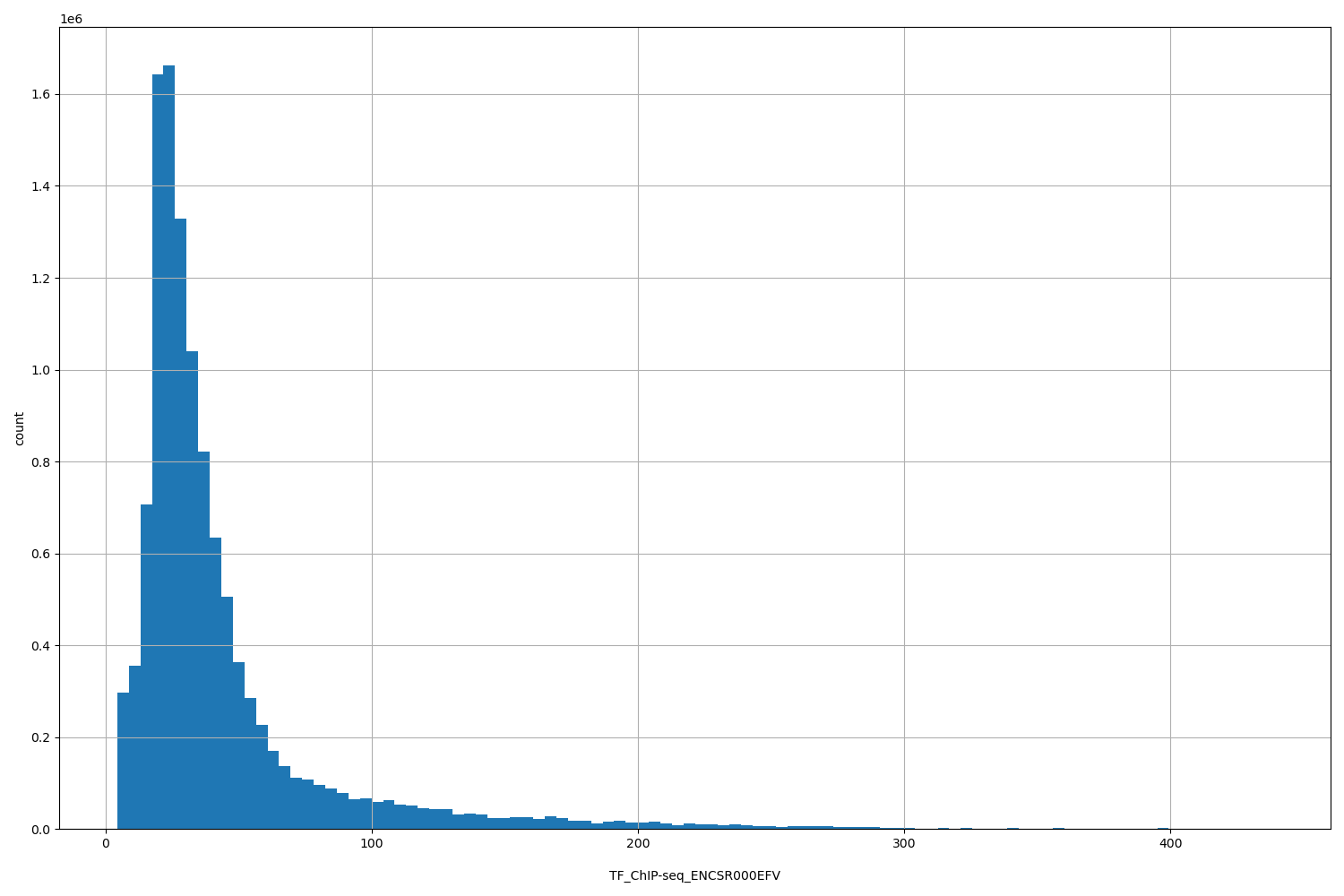
TF_ChIP-seq_ENCSR000EFV
| Id: | TF_ChIP-seq/ENCSR000EFV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EFV [biosamplesummary="Homo sapiens K562" and target="MAX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN985CVH|/analyses/ENCAN985CVH/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF618VMC|/files/ENCFF618VMC/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_not_compliant: Processed alignments file {ENCFF635HIS|/files/ENCFF635HIS/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 7403939 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF360QBV|/files/ENCFF360QBV/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 17947161 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF466VUY|/files/ENCFF466VUY/}, {ENCFF778PTE|/files/ENCFF778PTE/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 3.57. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF618VMC|/files/ENCFF618VMC/}, {ENCFF481AOS|/files/ENCFF481AOS/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 3.57. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EFV | float |
TF_ChIP-seq_ENCSR000EFV |
TF_ChIP-seq ENCSR000EFV [biosample_summary="Homo sapiens K562" and target="MAX"]
|
 |
[4.34, 438] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF618VMC.bed.gz | 610.95 KB | f4682a1dc246195895e23f28ea0ee326 |
| ENCFF618VMC.bed.gz.dvc | 100.0 B | 1596953fa599178e8cd6c8881b6953bd |
| ENCFF618VMC.tabix.bed.gz | 472.07 KB | d6c100511f125d10593219ffcd6988fc |
| ENCFF618VMC.tabix.bed.gz.dvc | 106.0 B | b36b271beb2e3f7c422a044695cb2a1f |
| ENCFF618VMC.tabix.bed.gz.tbi | 190.14 KB | b06bc5e068330bc6e38cdbd531d992d4 |
| ENCFF618VMC.tabix.bed.gz.tbi.dvc | 110.0 B | dcce2064699ed18bb88036ffa29b5265 |
| genomic_resource.yaml | 3.65 KB | abb9a070fcb158bf7518f3b3ec683e1f |
| genomic_resource_original.yaml | 3.56 KB | 304dd202171d9837042d3a8d86e7556b |
| statistics/ |