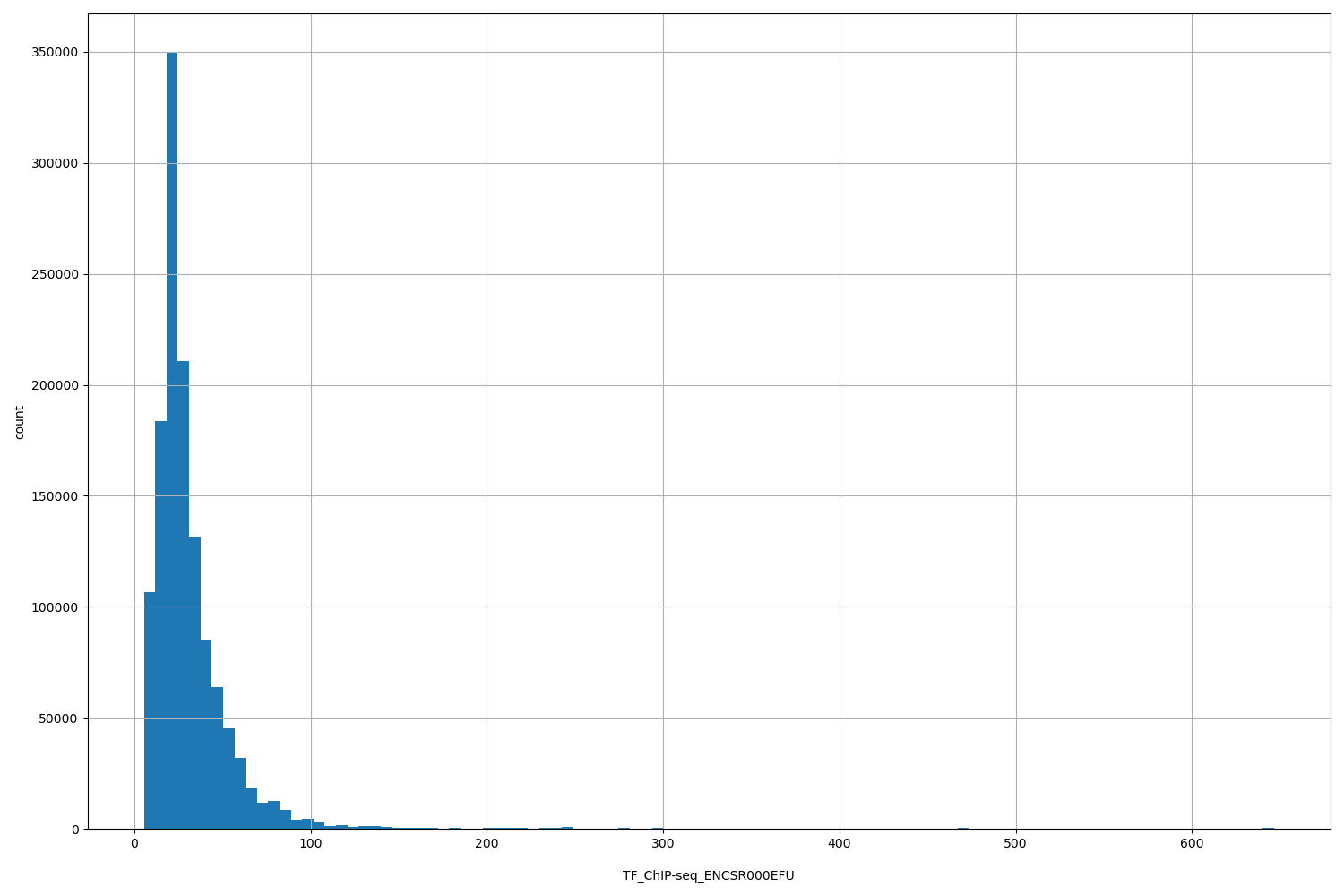
TF_ChIP-seq_ENCSR000EFU
| Id: | TF_ChIP-seq/ENCSR000EFU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EFU [biosamplesummary="Homo sapiens K562" and target="ELK1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN966RKN|/analyses/ENCAN966RKN/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF119SCQ|/files/ENCFF119SCQ/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: Processed alignments file {ENCFF865ISL|/files/ENCFF865ISL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 13948677 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ELK1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF554BLC|/files/ENCFF554BLC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 19075626 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ELK1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EFU | float |
TF_ChIP-seq_ENCSR000EFU |
TF_ChIP-seq ENCSR000EFU [biosample_summary="Homo sapiens K562" and target="ELK1"]
|
 |
[5.5, 647] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF119SCQ.bed.gz | 71.26 KB | d5791bd9656128411f3090b76911951b |
| ENCFF119SCQ.bed.gz.dvc | 99.0 B | 52ea7a18ceb84128d75e09155d9921d3 |
| ENCFF119SCQ.tabix.bed.gz | 51.56 KB | aa749c16f8dd26740df72ba82ac4de8f |
| ENCFF119SCQ.tabix.bed.gz.dvc | 105.0 B | ad2f797ed8354866eded0d93fb189fbe |
| ENCFF119SCQ.tabix.bed.gz.tbi | 36.91 KB | 8f054d576bd72da6709410a76108ab99 |
| ENCFF119SCQ.tabix.bed.gz.tbi.dvc | 109.0 B | dc6fe06efc2bdac9e5e0e471881184f9 |
| genomic_resource.yaml | 2.51 KB | 81f1bbbdf4af9ca80115d99615d97f51 |
| genomic_resource_original.yaml | 2.42 KB | 24ab1b5e95c7974a375d15e4f814b39f |
| statistics/ |