TF_ChIP-seq_ENCSR000EFB
| Id: | TF_ChIP-seq/ENCSR000EFB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EFB [biosamplesummary="Homo sapiens endothelial cell of umbilical vein newborn" and target="POLR2A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: newborn output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN496LMP|/analyses/ENCAN496LMP/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN496LMP|/analyses/ENCAN496LMP/} has in progress subobject document {59330728-be29-4816-9b9f-0ef62d49ecf9|/documents/59330728-be29-4816-9b9f-0ef62d49ecf9/} audit_not_compliant: Processed alignments file {ENCFF960EIT|/files/ENCFF960EIT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 7809067 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF362NDM|/files/ENCFF362NDM/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 8293447 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF561KEL|/files/ENCFF561KEL/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 7271235 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
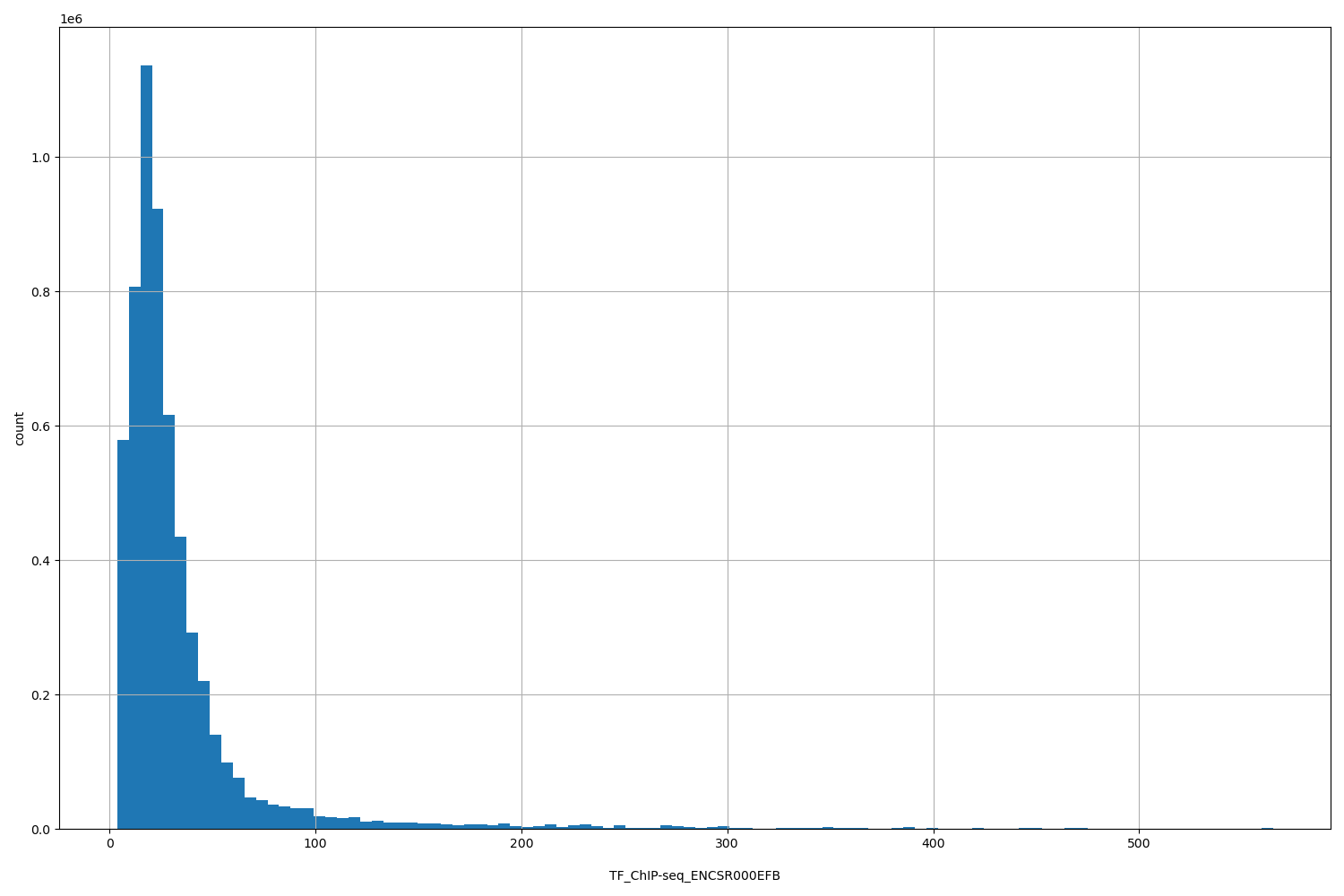
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EFB | float |
TF_ChIP-seq_ENCSR000EFB |
TF_ChIP-seq ENCSR000EFB [biosample_summary="Homo sapiens endothelial cell of umbilical vein newborn" and target="POLR2A"]
|
 |
[3.82, 565] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF870KJE.bed.gz | 287.89 KB | b6c0a2b1238f12e8f7b303636f7fd62b |
| ENCFF870KJE.bed.gz.dvc | 100.0 B | 16db7535ad17e221422e0b47f474d44c |
| ENCFF870KJE.tabix.bed.gz | 225.57 KB | 7d11ee86d4bbb61d7cca667869002cac |
| ENCFF870KJE.tabix.bed.gz.dvc | 106.0 B | 63947c7509d550cb0b25b42ae9149120 |
| ENCFF870KJE.tabix.bed.gz.tbi | 88.79 KB | 94bc68a04678179c71498dd6a86ee111 |
| ENCFF870KJE.tabix.bed.gz.tbi.dvc | 109.0 B | 9d24f5512b67630bcf35a0583b57d639 |
| genomic_resource.yaml | 3.31 KB | e797af05d61d643a4f419c19146aea3b |
| genomic_resource_original.yaml | 3.18 KB | 2d212e452714756d39d4a0ac42938839 |
| statistics/ |