TF_ChIP-seq_ENCSR000EEZ
| Id: | TF_ChIP-seq/ENCSR000EEZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EEZ [biosamplesummary="Homo sapiens endothelial cell of umbilical vein newborn" and target="MAX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: newborn output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN114NRZ|/analyses/ENCAN114NRZ/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN114NRZ|/analyses/ENCAN114NRZ/} has in progress subobject document {6f779aba-4605-4410-a819-0c80d0e646c2|/documents/6f779aba-4605-4410-a819-0c80d0e646c2/} audit_not_compliant: Processed alignments file {ENCFF801OVR|/files/ENCFF801OVR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9335346 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF302DYJ|/files/ENCFF302DYJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6983721 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
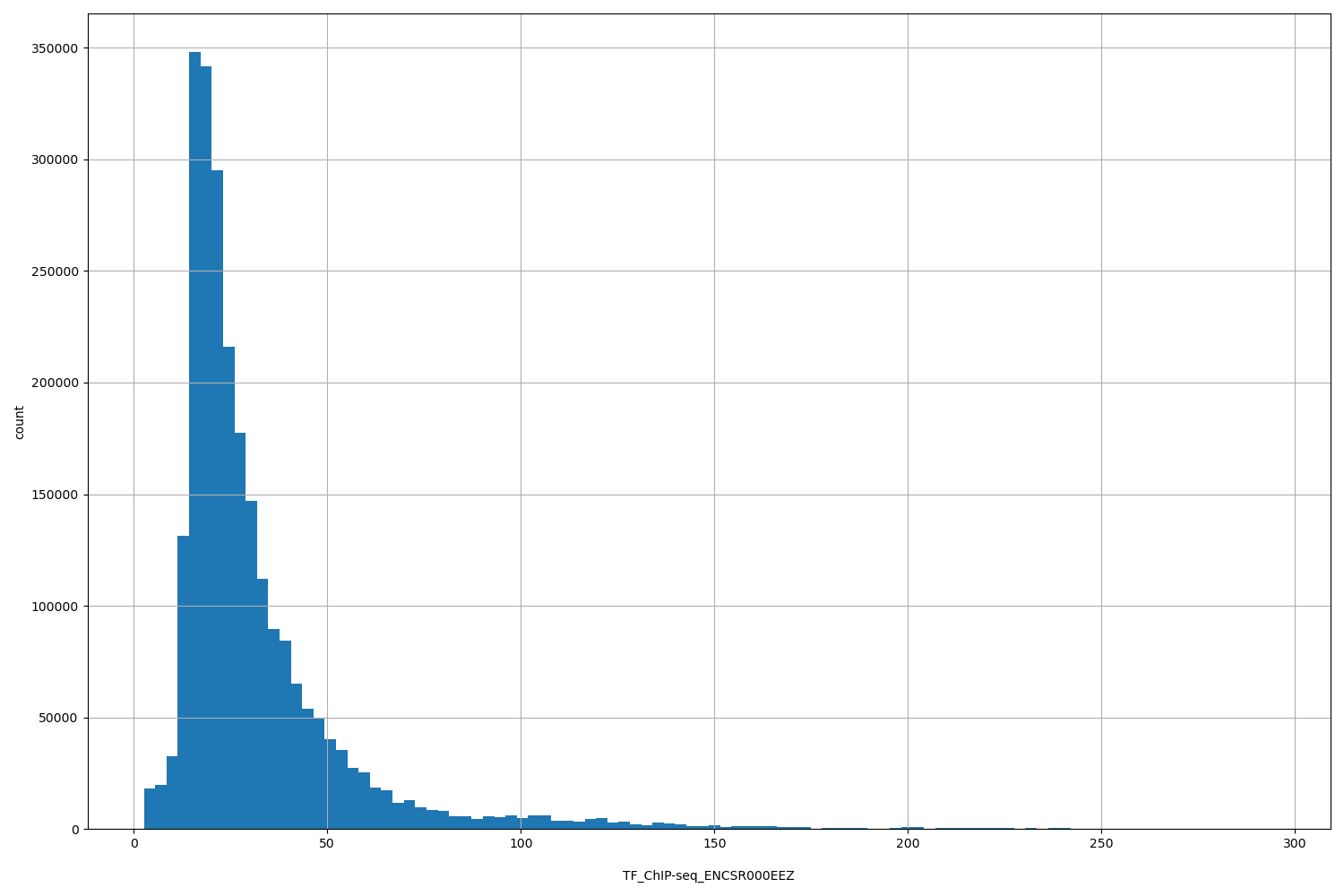
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EEZ | float |
TF_ChIP-seq_ENCSR000EEZ |
TF_ChIP-seq ENCSR000EEZ [biosample_summary="Homo sapiens endothelial cell of umbilical vein newborn" and target="MAX"]
|
 |
[2.63, 295] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF562ZOV.bed.gz | 186.68 KB | b5fe5760382c2e6cc6c228644b54fe08 |
| ENCFF562ZOV.bed.gz.dvc | 100.0 B | 6720c55bebf3c01364455c37b43f5565 |
| ENCFF562ZOV.tabix.bed.gz | 130.61 KB | e4eadba37a06530cb12c06646862b9a8 |
| ENCFF562ZOV.tabix.bed.gz.dvc | 106.0 B | c62de1249da859d90971dada6b65331f |
| ENCFF562ZOV.tabix.bed.gz.tbi | 89.05 KB | 7fc489a4fcb2a133d13f0493ab2942b5 |
| ENCFF562ZOV.tabix.bed.gz.tbi.dvc | 109.0 B | 426230a5520580bcbc46f3b36fefd614 |
| genomic_resource.yaml | 2.8 KB | 837dc11311d45dea6b32d378fefe9e19 |
| genomic_resource_original.yaml | 2.67 KB | 8de6e43b238196efcfd3542b4c18b47e |
| statistics/ |