TF_ChIP-seq_ENCSR000EEF
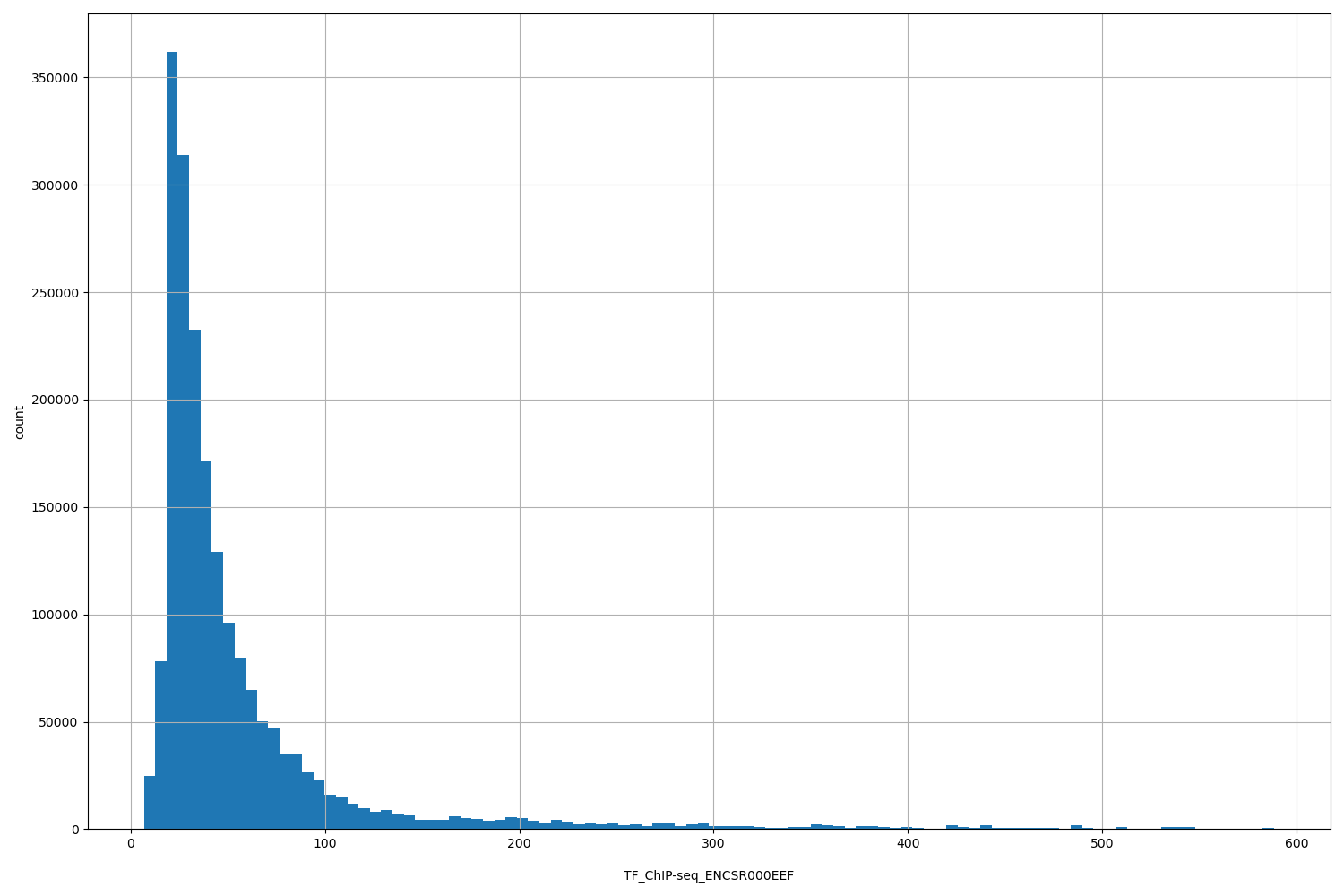
| Id: | TF_ChIP-seq/ENCSR000EEF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EEF [biosamplesummary="Homo sapiens HepG2" and target="USF2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN932RXZ|/analyses/ENCAN932RXZ/} has in progress subobject document {fdd87abb-4968-4a9a-b0cd-f7140a064eea|/documents/fdd87abb-4968-4a9a-b0cd-f7140a064eea/} audit_internal_action: Released analysis {ENCAN932RXZ|/analyses/ENCAN932RXZ/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF787PWA|/files/ENCFF787PWA/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 17185936 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting USF2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EEF | float |
TF_ChIP-seq_ENCSR000EEF |
TF_ChIP-seq ENCSR000EEF [biosample_summary="Homo sapiens HepG2" and target="USF2"]
|
 |
[6.7, 588] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF652ELW.bed.gz | 142.14 KB | 3faf4a3f841372df7e9cd763d30d4e06 |
| ENCFF652ELW.bed.gz.dvc | 100.0 B | c6274a259a176782edd6700d82faad25 |
| ENCFF652ELW.tabix.bed.gz | 95.64 KB | d335961305460a886a4857bda176c7de |
| ENCFF652ELW.tabix.bed.gz.dvc | 105.0 B | 044fb0278cb87217d5087ff87fb38732 |
| ENCFF652ELW.tabix.bed.gz.tbi | 71.87 KB | 10718b4814caa2bfed77bc524759c580 |
| ENCFF652ELW.tabix.bed.gz.tbi.dvc | 109.0 B | cfabeab16e1622ccb8988eace04115e1 |
| genomic_resource.yaml | 2.05 KB | ea7e5567dc6eca925fd6c1b28974e571 |
| genomic_resource_original.yaml | 1.96 KB | e5b196a0374d305942ead3da99afc100 |
| statistics/ |