TF_ChIP-seq_ENCSR000EDP
| Id: | TF_ChIP-seq/ENCSR000EDP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDP [biosamplesummary="Homo sapiens HepG2" and target="ARID3A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN767KUT|/analyses/ENCAN767KUT/} has in progress subobject document {8735b908-0695-4e36-ba32-df99732da089|/documents/8735b908-0695-4e36-ba32-df99732da089/} audit_internal_action: Released analysis {ENCAN767KUT|/analyses/ENCAN767KUT/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF432MNS|/files/ENCFF432MNS/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19044340 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ARID3A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF900KAB|/files/ENCFF900KAB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18729112 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ARID3A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
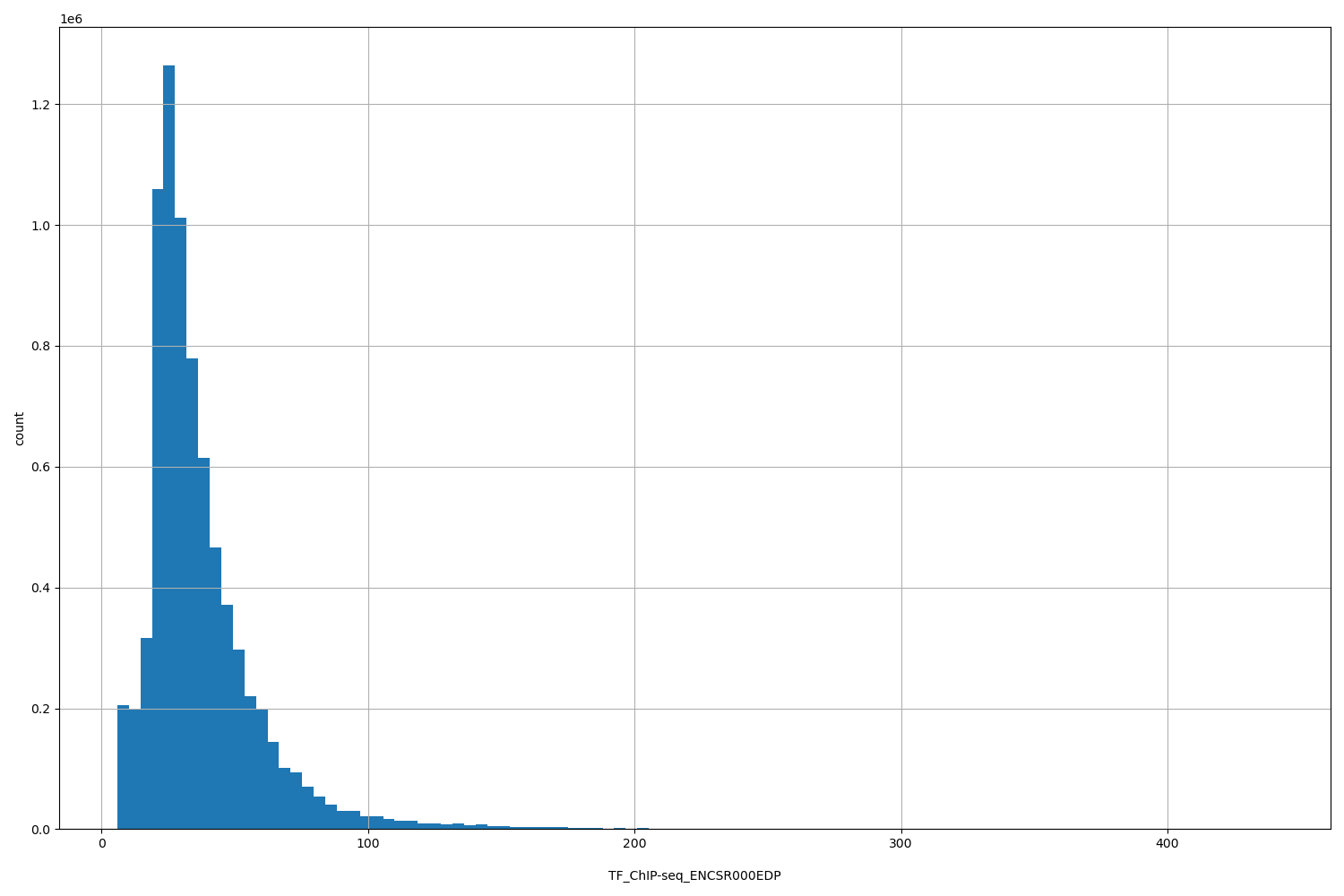
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDP | float |
TF_ChIP-seq_ENCSR000EDP |
TF_ChIP-seq ENCSR000EDP [biosample_summary="Homo sapiens HepG2" and target="ARID3A"]
|
 |
[5.9, 440] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF536DJU.bed.gz | 395.15 KB | f045c4efc21c1584e18d91942ab5f8d3 |
| ENCFF536DJU.bed.gz.dvc | 100.0 B | dddbabbe59ec503b6ec36f0e4b40ff45 |
| ENCFF536DJU.tabix.bed.gz | 274.03 KB | 753400e596398fb46fb3ded4660a50b6 |
| ENCFF536DJU.tabix.bed.gz.dvc | 106.0 B | ea19391c2060279c9e8e0dec4194b186 |
| ENCFF536DJU.tabix.bed.gz.tbi | 143.59 KB | 10e07b14b8e2ccf5bb4ef55461b8e761 |
| ENCFF536DJU.tabix.bed.gz.tbi.dvc | 110.0 B | ab8baeacde45af0cc06c21e9b17096ec |
| genomic_resource.yaml | 2.57 KB | 42a44e1d9c2254b9dd71cf3ac3d00f8f |
| genomic_resource_original.yaml | 2.47 KB | 4a8e178f30961e77b603ab2f802ae8ac |
| statistics/ |