TF_ChIP-seq_ENCSR000EDL
| Id: | TF_ChIP-seq/ENCSR000EDL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDL [biosamplesummary="Homo sapiens HeLa-S3" and target="SMARCC2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2, Rep 3 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN592WYV|/analyses/ENCAN592WYV/} has in progress subobject document {6fe05b58-db81-463a-a352-f9cf98c566fb|/documents/6fe05b58-db81-463a-a352-f9cf98c566fb/} audit_internal_action: Released analysis {ENCAN592WYV|/analyses/ENCAN592WYV/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF485JQN|/files/ENCFF485JQN/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9534665 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SMARCC2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF439SBY|/files/ENCFF439SBY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9440995 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SMARCC2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF022SZY|/files/ENCFF022SZY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9746381 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SMARCC2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
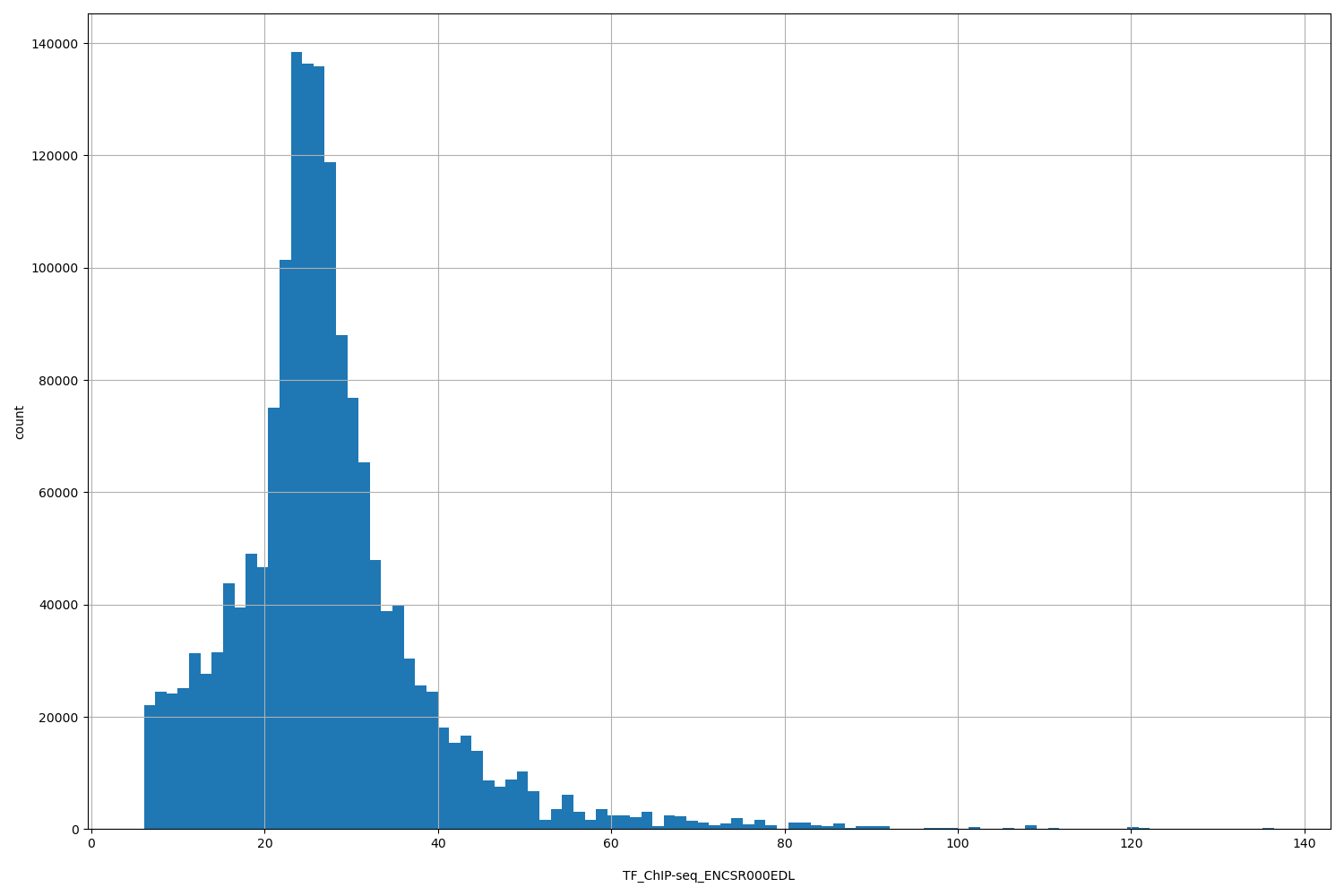
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDL | float |
TF_ChIP-seq_ENCSR000EDL |
TF_ChIP-seq ENCSR000EDL [biosample_summary="Homo sapiens HeLa-S3" and target="SMARCC2"]
|
 |
[6.06, 137] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF253UAA.bed.gz | 71.98 KB | 9e2e8f95e91a1945c624b08921e6e054 |
| ENCFF253UAA.bed.gz.dvc | 99.0 B | baf1367f51baf85c787f49c215e8fc27 |
| ENCFF253UAA.tabix.bed.gz | 51.91 KB | b826bc42631f40ce3d57cfd6bd354add |
| ENCFF253UAA.tabix.bed.gz.dvc | 105.0 B | 839a77f41965c04569892fa3586772f5 |
| ENCFF253UAA.tabix.bed.gz.tbi | 33.84 KB | b8de461195c811937f50a52aa97aace5 |
| ENCFF253UAA.tabix.bed.gz.tbi.dvc | 109.0 B | 3617ca72f057248e1c52dbf5bdd37ade |
| genomic_resource.yaml | 3.11 KB | 6b54e5113f498ff3c0298c679ae098a4 |
| genomic_resource_original.yaml | 3.01 KB | ac659f3ad0e7ad4d79c23fa84cba0a62 |
| statistics/ |