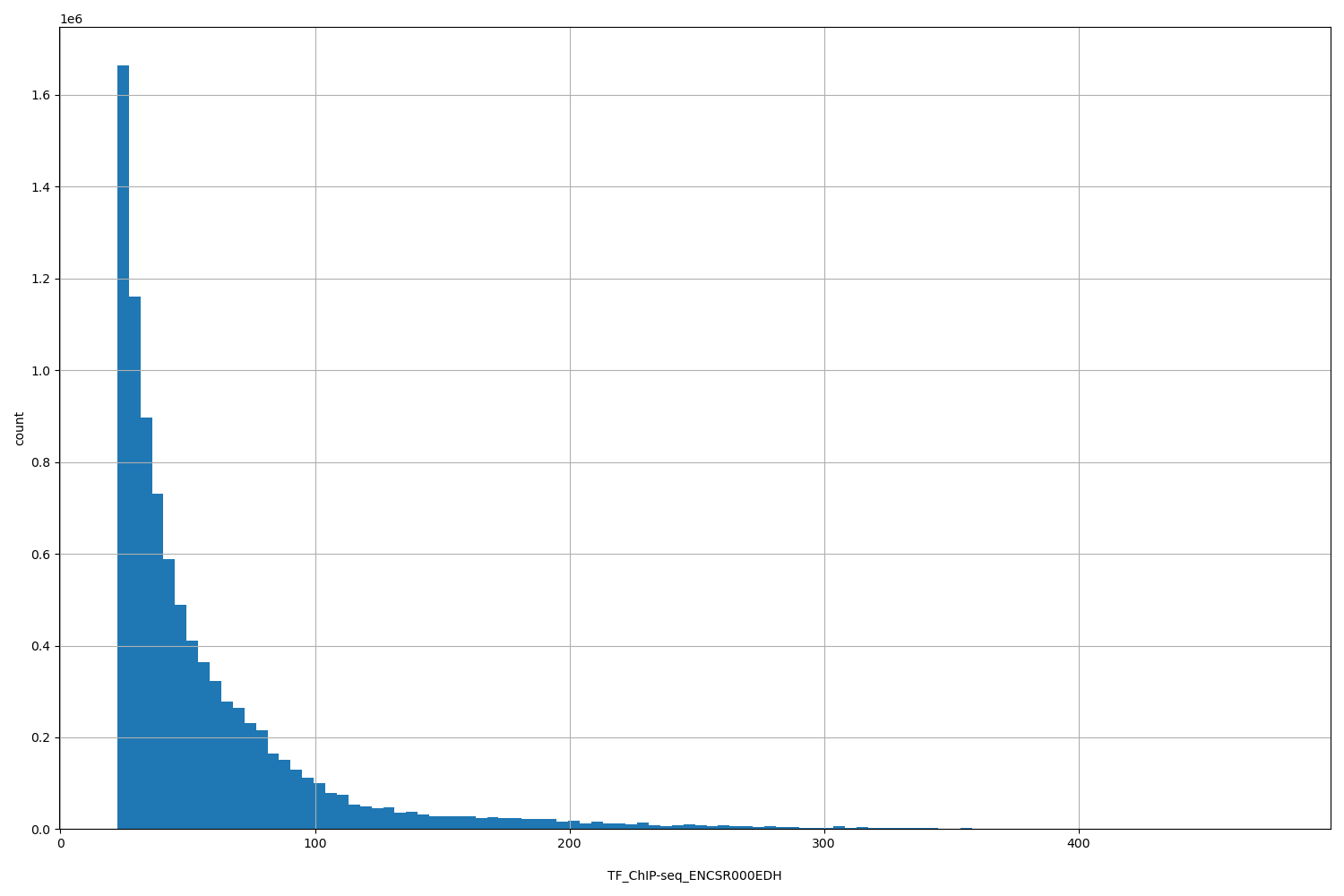
TF_ChIP-seq_ENCSR000EDH
TF_ChIP-seq ENCSR000EDH [biosample_summary="Homo sapiens HeLa-S3" and target="JUND"]
| Id: | TF_ChIP-seq/ENCSR000EDH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDH [biosamplesummary="Homo sapiens HeLa-S3" and target="JUND"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CSP|/files/ENCFF002CSP/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDH | float |
TF_ChIP-seq_ENCSR000EDH |
TF_ChIP-seq ENCSR000EDH [biosample_summary="Homo sapiens HeLa-S3" and target="JUND"]
|
 |
[22.3, 476] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CSP.bed.gz | 733.0 KB | 8382b1d2abe95d66c1230cf762e7e99e |
| ENCFF002CSP.bed.gz.dvc | 100.0 B | 0992ba35ec6e5820c280a963201a67d8 |
| ENCFF002CSP.tabix.bed.gz | 482.87 KB | 50e2688468927b60781cc094d4d0672d |
| ENCFF002CSP.tabix.bed.gz.dvc | 106.0 B | 7f6c08b7573833d5ddb0f931746a715b |
| ENCFF002CSP.tabix.bed.gz.tbi | 202.04 KB | 6360bb6e05ad36aa574e828accfbb270 |
| ENCFF002CSP.tabix.bed.gz.tbi.dvc | 110.0 B | 08d976cb6c34d96302baec9e63f005b8 |
| genomic_resource.yaml | 1.48 KB | 6b713140248dfe2c8bf414c506bb8adc |
| genomic_resource_original.yaml | 1.38 KB | 7e48a3c4280003fb56d3bb4fffb98d1a |
| statistics/ |