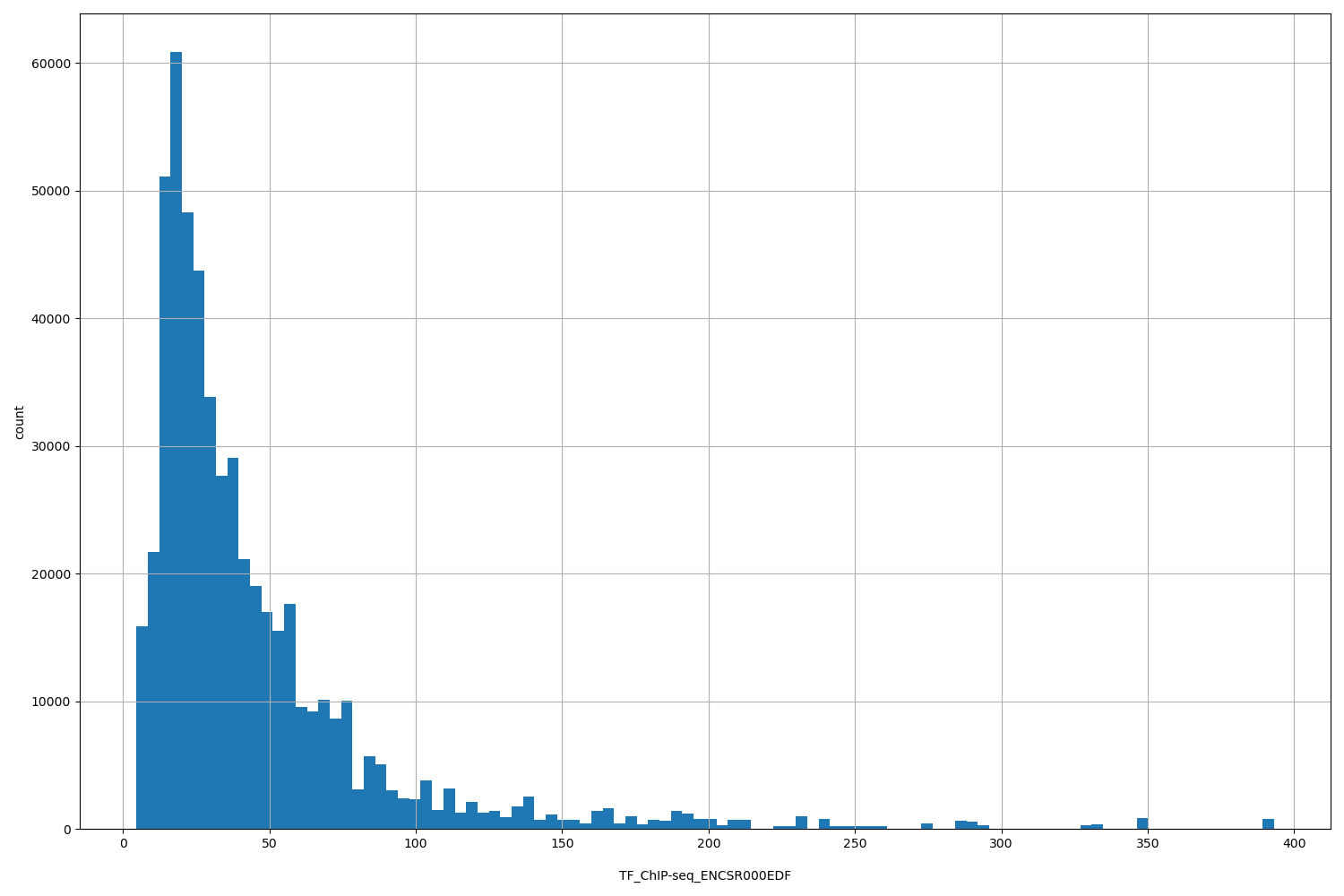
TF_ChIP-seq_ENCSR000EDF
| Id: | TF_ChIP-seq/ENCSR000EDF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDF [biosamplesummary="Homo sapiens HeLa-S3" and target="IRF3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN793KOB|/analyses/ENCAN793KOB/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN793KOB|/analyses/ENCAN793KOB/} has in progress subobject document {4a5eb352-43c1-402e-9b57-728bf47b4aaa|/documents/4a5eb352-43c1-402e-9b57-728bf47b4aaa/} audit_warning: Processed alignments file {ENCFF701FQQ|/files/ENCFF701FQQ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13025860 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting IRF3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF873UFD|/files/ENCFF873UFD/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13665420 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting IRF3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDF | float |
TF_ChIP-seq_ENCSR000EDF |
TF_ChIP-seq ENCSR000EDF [biosample_summary="Homo sapiens HeLa-S3" and target="IRF3"]
|
 |
[4.57, 393] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF291VFT.bed.gz | 34.21 KB | 5e4d1196d4106a38009215c10578156e |
| ENCFF291VFT.bed.gz.dvc | 99.0 B | 0b41d06db89041ba293f38bdf8a44886 |
| ENCFF291VFT.tabix.bed.gz | 23.26 KB | d1fe94e4bbe59c8090a37de6c48fba4b |
| ENCFF291VFT.tabix.bed.gz.dvc | 105.0 B | 294673e155309e5bd94fa44590bffd1b |
| ENCFF291VFT.tabix.bed.gz.tbi | 22.32 KB | be2c517c712bd1f2864d4ae344b8ab93 |
| ENCFF291VFT.tabix.bed.gz.tbi.dvc | 109.0 B | 032760f5c3ff970165656114d6788776 |
| genomic_resource.yaml | 2.56 KB | d82ce01d6b970780d73c4e7634110cb5 |
| genomic_resource_original.yaml | 2.47 KB | d1e929ca27d456914590deb716221d28 |
| statistics/ |