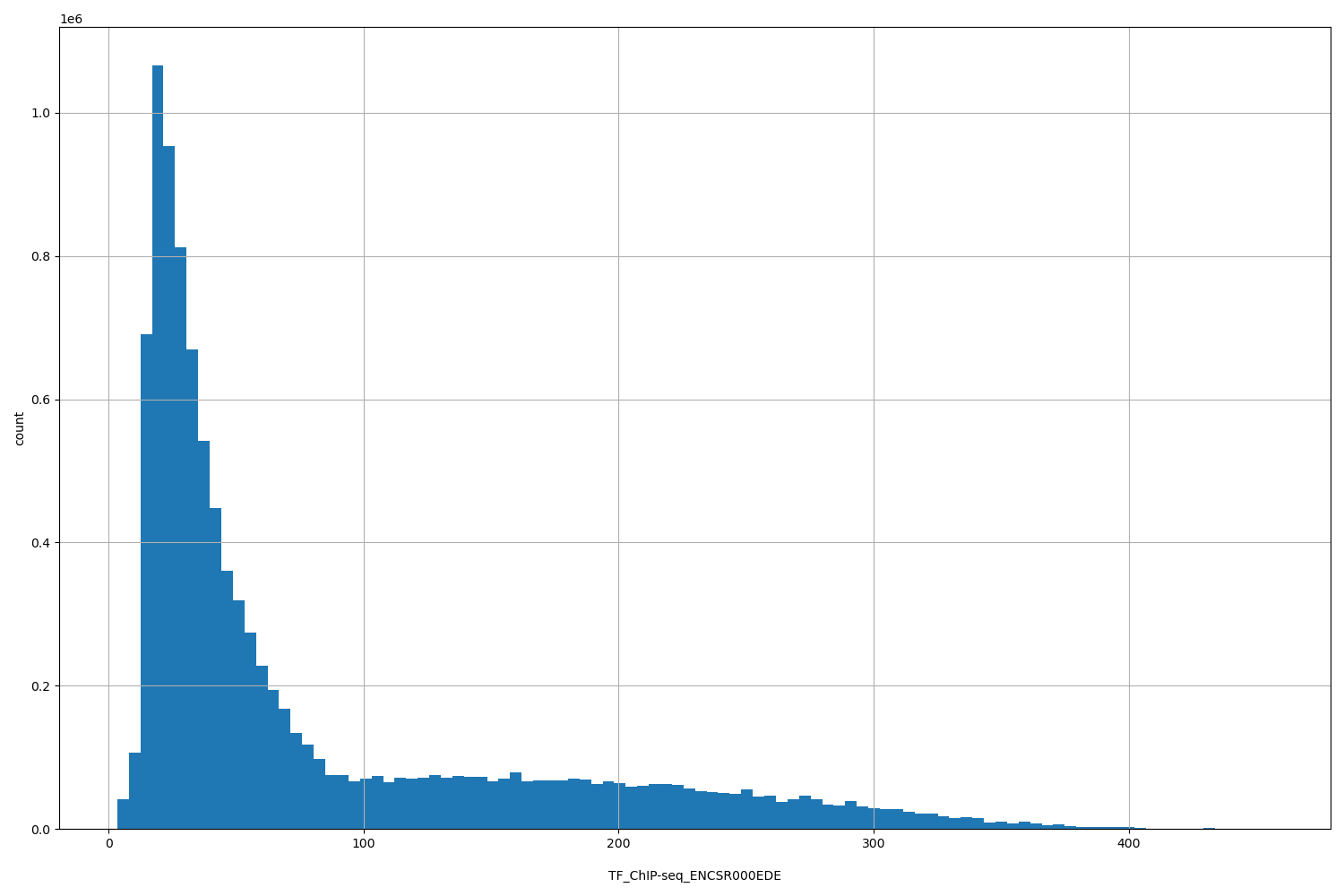
TF_ChIP-seq_ENCSR000EDE
| Id: | TF_ChIP-seq/ENCSR000EDE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDE [biosamplesummary="Homo sapiens HeLa-S3" and target="RAD21"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN118XQA|/analyses/ENCAN118XQA/} has in progress subobject document {364cd64d-91ec-43ec-99be-0c10efab75ec|/documents/364cd64d-91ec-43ec-99be-0c10efab75ec/} audit_internal_action: Released analysis {ENCAN118XQA|/analyses/ENCAN118XQA/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF705SCW|/files/ENCFF705SCW/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 12129259 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting RAD21-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF632OBR|/files/ENCFF632OBR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13139407 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting RAD21-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDE | float |
TF_ChIP-seq_ENCSR000EDE |
TF_ChIP-seq ENCSR000EDE [biosample_summary="Homo sapiens HeLa-S3" and target="RAD21"]
|
 |
[3.44, 456] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF239FBO.bed.gz | 843.44 KB | e4237b0767796723b4a7a51f0d6c0463 |
| ENCFF239FBO.bed.gz.dvc | 100.0 B | 40ab5dccd99a9297d5974ef61fe06ffe |
| ENCFF239FBO.tabix.bed.gz | 595.17 KB | a01d66ca8e0096e08cf0ee5892a0600e |
| ENCFF239FBO.tabix.bed.gz.dvc | 106.0 B | b124b009721b17c13ec78012d2fffb3a |
| ENCFF239FBO.tabix.bed.gz.tbi | 307.02 KB | a6aa258bede75339b514c68718a1326b |
| ENCFF239FBO.tabix.bed.gz.tbi.dvc | 110.0 B | 54100b483182226f75f33952a86d1a6e |
| genomic_resource.yaml | 2.57 KB | 3d4fae994c1ebf3f9c2647cdfb5a5ff1 |
| genomic_resource_original.yaml | 2.47 KB | 0405f39811d048106f98b2e518ee1100 |
| statistics/ |