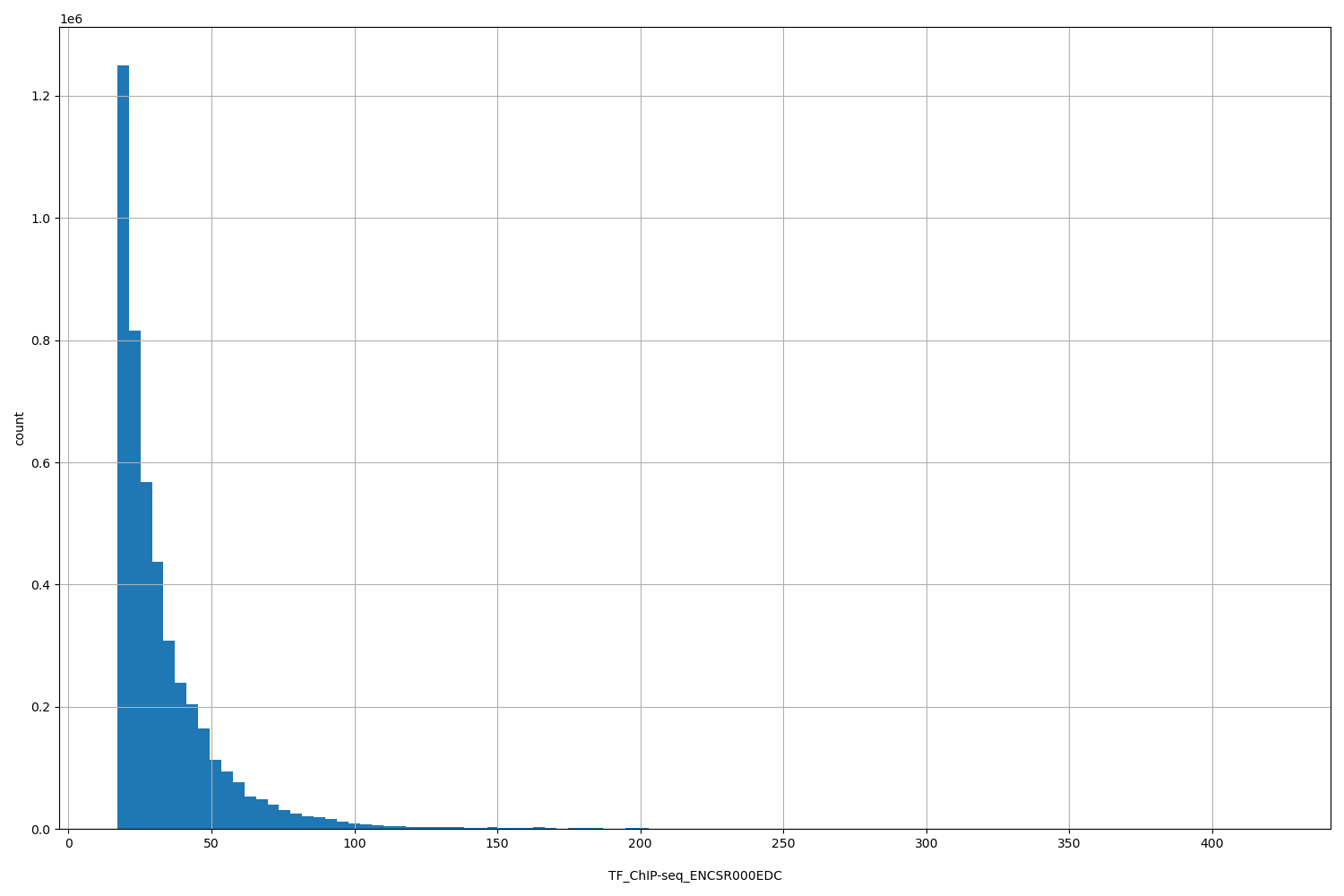
TF_ChIP-seq_ENCSR000EDC
TF_ChIP-seq ENCSR000EDC [biosample_summary="Homo sapiens HeLa-S3" and target="STAT3"]
| Id: | TF_ChIP-seq/ENCSR000EDC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EDC [biosamplesummary="Homo sapiens HeLa-S3" and target="STAT3"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CTH|/files/ENCFF002CTH/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EDC | float |
TF_ChIP-seq_ENCSR000EDC |
TF_ChIP-seq ENCSR000EDC [biosample_summary="Homo sapiens HeLa-S3" and target="STAT3"]
|
 |
[17.1, 421] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CTH.bed.gz | 340.36 KB | 9d04aa699bf6a49606986f1798d9d4f8 |
| ENCFF002CTH.bed.gz.dvc | 100.0 B | e465ebd5983e81ca2fe98e7c506c46b7 |
| ENCFF002CTH.tabix.bed.gz | 213.05 KB | 5aeff634157584c5f0e5ea0d4a4f89fb |
| ENCFF002CTH.tabix.bed.gz.dvc | 106.0 B | 6fcec605ca10ec29bf1241c43a82fa94 |
| ENCFF002CTH.tabix.bed.gz.tbi | 102.81 KB | ca561baef31ac8f9bf6479e09d313a7e |
| ENCFF002CTH.tabix.bed.gz.tbi.dvc | 110.0 B | b1e1dd35c0bb506207559df3c4a992d1 |
| genomic_resource.yaml | 1.48 KB | 06587c08c5bfd475f5683630bfda2853 |
| genomic_resource_original.yaml | 1.38 KB | 520b0628f5cca9bc37d8e97bff1d7c3c |
| statistics/ |