TF_ChIP-seq_ENCSR000ECS
TF_ChIP-seq ENCSR000ECS [biosample_summary="Homo sapiens HeLa-S3" and target="SMC3"]
| Id: | TF_ChIP-seq/ENCSR000ECS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000ECS [biosamplesummary="Homo sapiens HeLa-S3" and target="SMC3"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CTE|/files/ENCFF002CTE/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
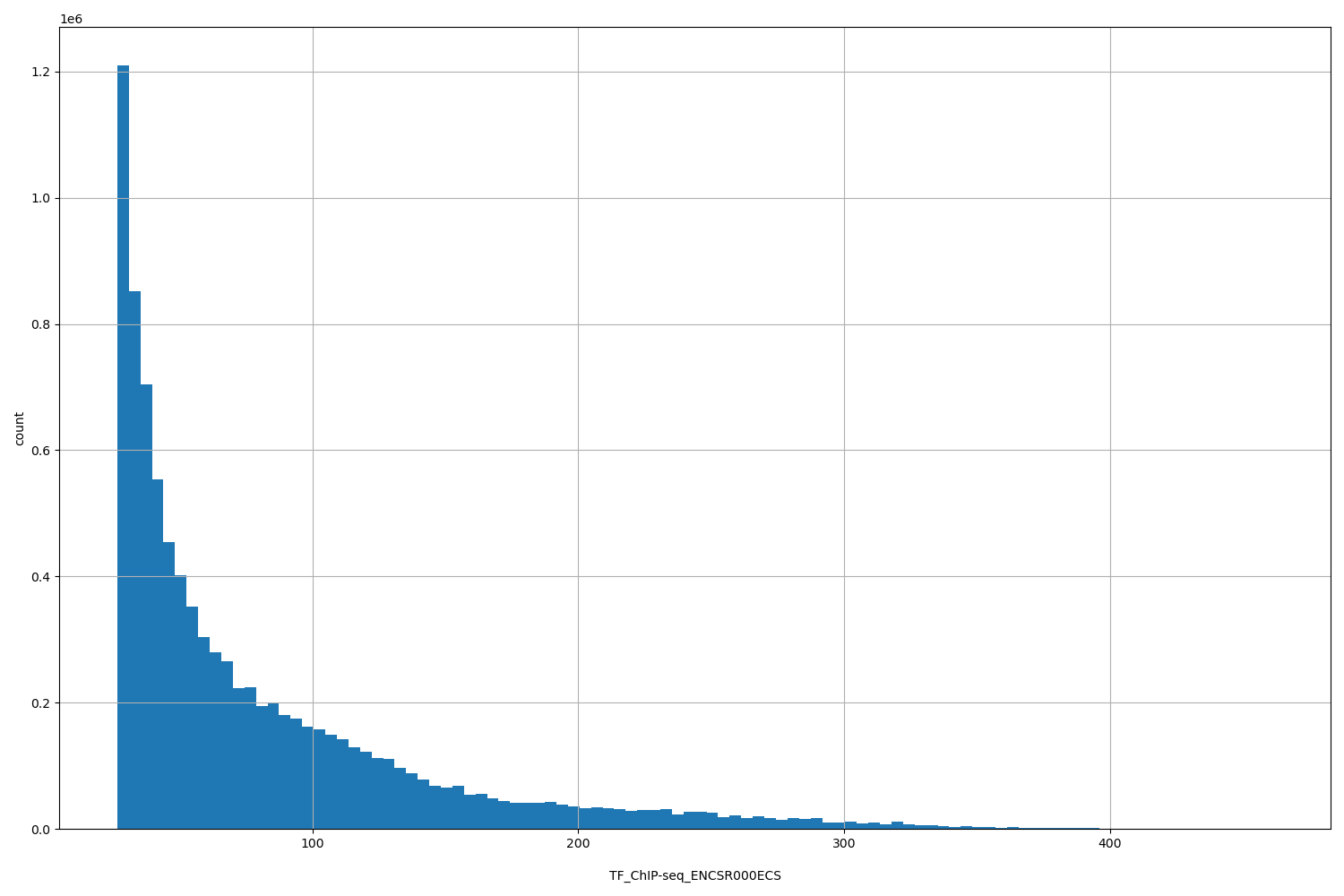
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000ECS | float |
TF_ChIP-seq_ENCSR000ECS |
TF_ChIP-seq ENCSR000ECS [biosample_summary="Homo sapiens HeLa-S3" and target="SMC3"]
|
 |
[26.6, 461] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CTE.bed.gz | 917.46 KB | 4fbd9b20cc5b419c3086e3ca689fe9ee |
| ENCFF002CTE.bed.gz.dvc | 100.0 B | 67636f29747cdbd953935eaff710eecd |
| ENCFF002CTE.tabix.bed.gz | 600.56 KB | 3ee9a97af0a0ede36ad5d7eb5eaa84e3 |
| ENCFF002CTE.tabix.bed.gz.dvc | 106.0 B | 1268c7ca23131841c04d4890c78fd538 |
| ENCFF002CTE.tabix.bed.gz.tbi | 253.06 KB | 9d2941096c8bc1c29f167ee3eace4503 |
| ENCFF002CTE.tabix.bed.gz.tbi.dvc | 110.0 B | dc1b9ea051ebf6ec1506372768cd576b |
| genomic_resource.yaml | 1.48 KB | c74a15ae2f6797e6a0a60b725e00ce17 |
| genomic_resource_original.yaml | 1.38 KB | c1d7edc368f2fc3ed533ab26cea06d7c |
| statistics/ |