TF_ChIP-seq_ENCSR000ECP
TF_ChIP-seq ENCSR000ECP [biosample_summary="Homo sapiens HeLa-S3" and target="CHD2"]
| Id: | TF_ChIP-seq/ENCSR000ECP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000ECP [biosamplesummary="Homo sapiens HeLa-S3" and target="CHD2"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CSC|/files/ENCFF002CSC/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
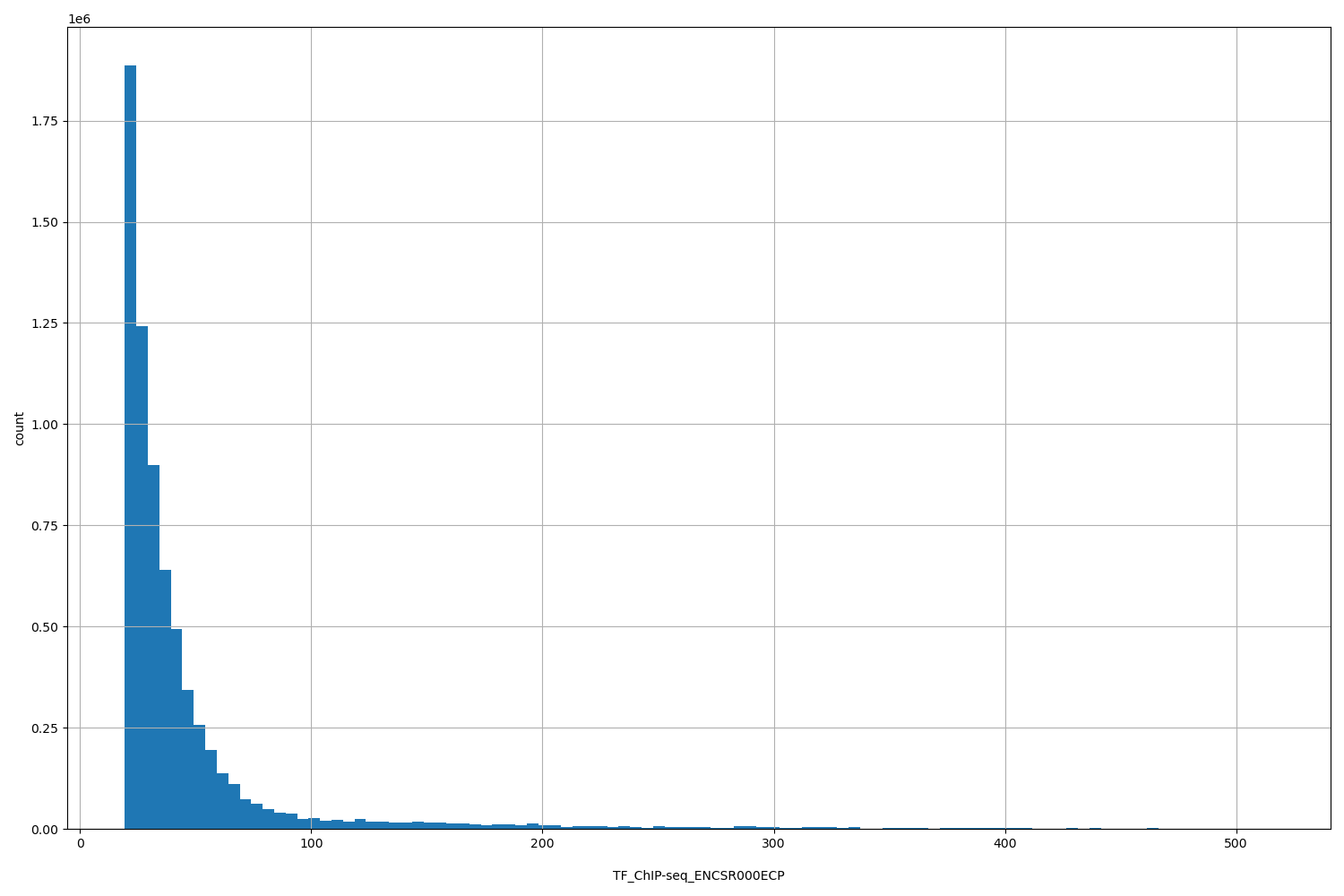
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000ECP | float |
TF_ChIP-seq_ENCSR000ECP |
TF_ChIP-seq ENCSR000ECP [biosample_summary="Homo sapiens HeLa-S3" and target="CHD2"]
|
 |
[19.4, 516] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CSC.bed.gz | 497.81 KB | 74a3d8795885876e1e4b953d6b54fe98 |
| ENCFF002CSC.bed.gz.dvc | 100.0 B | 321b7b88f82b372262a5706dde2b7cb7 |
| ENCFF002CSC.tabix.bed.gz | 319.27 KB | aebb50ec85962bcd684df91617ce33de |
| ENCFF002CSC.tabix.bed.gz.dvc | 106.0 B | 3499593af7db89f01724762075664955 |
| ENCFF002CSC.tabix.bed.gz.tbi | 132.53 KB | 25cca58f5f16cdc47329de5508726151 |
| ENCFF002CSC.tabix.bed.gz.tbi.dvc | 110.0 B | 09bde88a6fa02adbca402aa8e339d1fa |
| genomic_resource.yaml | 1.48 KB | 4b567d99acf564794ff9cc2b9dbf0dee |
| genomic_resource_original.yaml | 1.38 KB | 3a554a6310f9c5ee1063228737993f64 |
| statistics/ |