TF_ChIP-seq_ENCSR000ECM
TF_ChIP-seq ENCSR000ECM [biosample_summary="Homo sapiens HeLa-S3" and target="RCOR1"]
| Id: | TF_ChIP-seq/ENCSR000ECM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000ECM [biosamplesummary="Homo sapiens HeLa-S3" and target="RCOR1"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CSF|/files/ENCFF002CSF/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
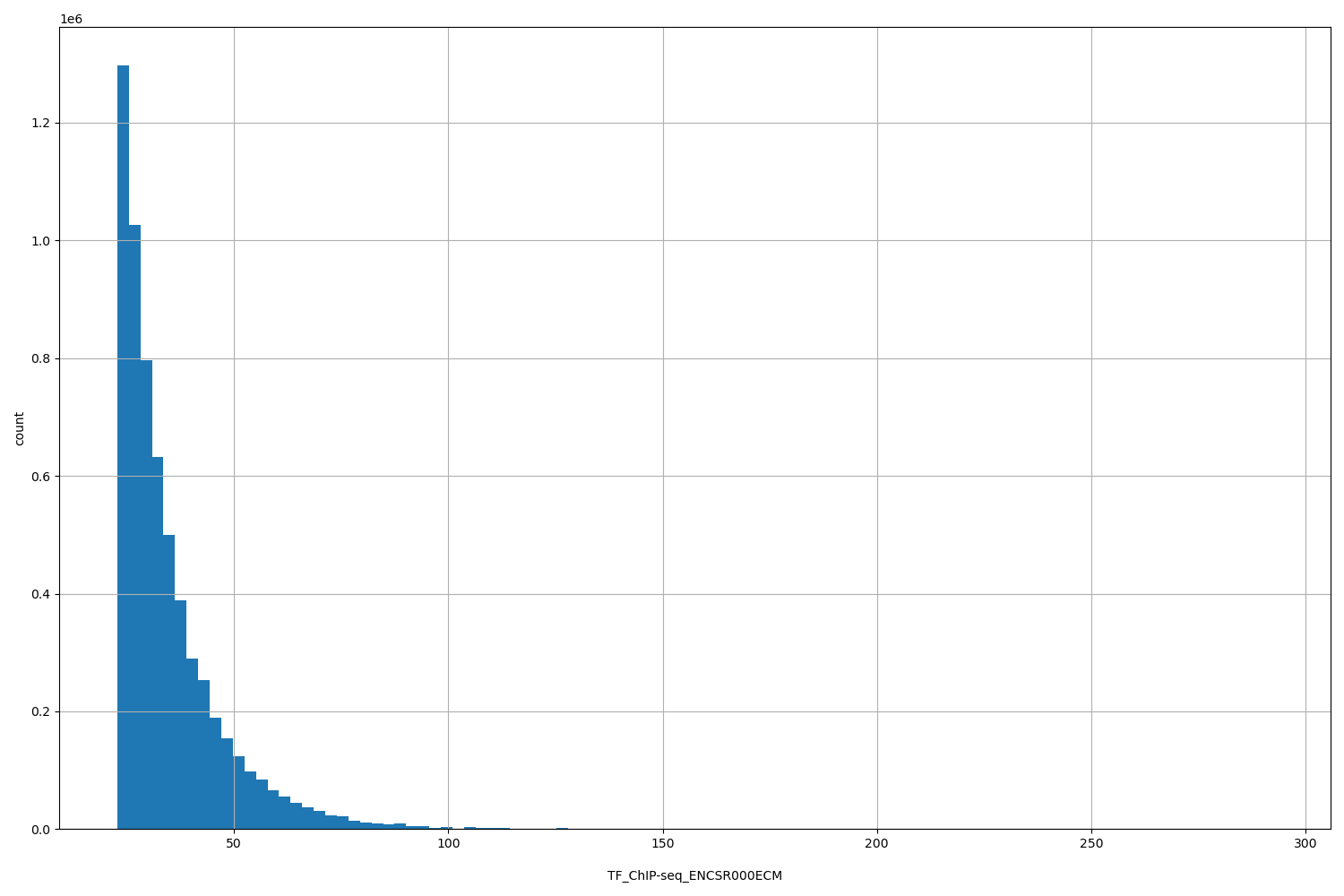
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000ECM | float |
TF_ChIP-seq_ENCSR000ECM |
TF_ChIP-seq ENCSR000ECM [biosample_summary="Homo sapiens HeLa-S3" and target="RCOR1"]
|
 |
[22.7, 292] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CSF.bed.gz | 385.63 KB | e7c99d8ef0836c01ae22ce43c8fbe510 |
| ENCFF002CSF.bed.gz.dvc | 100.0 B | 4c46f164a60d7c4f35eb86645f61c3b2 |
| ENCFF002CSF.tabix.bed.gz | 252.87 KB | 9919b186928b2cc7c47d427eda79f4e3 |
| ENCFF002CSF.tabix.bed.gz.dvc | 106.0 B | 56d5d7674fe3975dd4c88f6f51c5d7e0 |
| ENCFF002CSF.tabix.bed.gz.tbi | 116.79 KB | 6f1ea8367049f70232b2c852da5eb102 |
| ENCFF002CSF.tabix.bed.gz.tbi.dvc | 110.0 B | cf09054e77f62e63e54e26fcfd7562c8 |
| genomic_resource.yaml | 1.48 KB | 1db581b79fcff147e3b8dde25f87df68 |
| genomic_resource_original.yaml | 1.38 KB | 809ba25547f52593ba292dec49f2271f |
| statistics/ |