TF_ChIP-seq_ENCSR000EBP
| Id: | TF_ChIP-seq/ENCSR000EBP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EBP [biosamplesummary="Homo sapiens H1" and target="GTF2F1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN274BJE|/analyses/ENCAN274BJE/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF509UIP|/files/ENCFF509UIP/} processed by ChIP-seq ENCODE3 hg19 pipeline has 16361310 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting GTF2F1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF205YOD|/files/ENCFF205YOD/}, {ENCFF214III|/files/ENCFF214III/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.08 and a self consistency ratio of 2.05. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF750MMS|/files/ENCFF750MMS/}, {ENCFF922KXQ|/files/ENCFF922KXQ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.08 and a self consistency ratio of 2.05. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
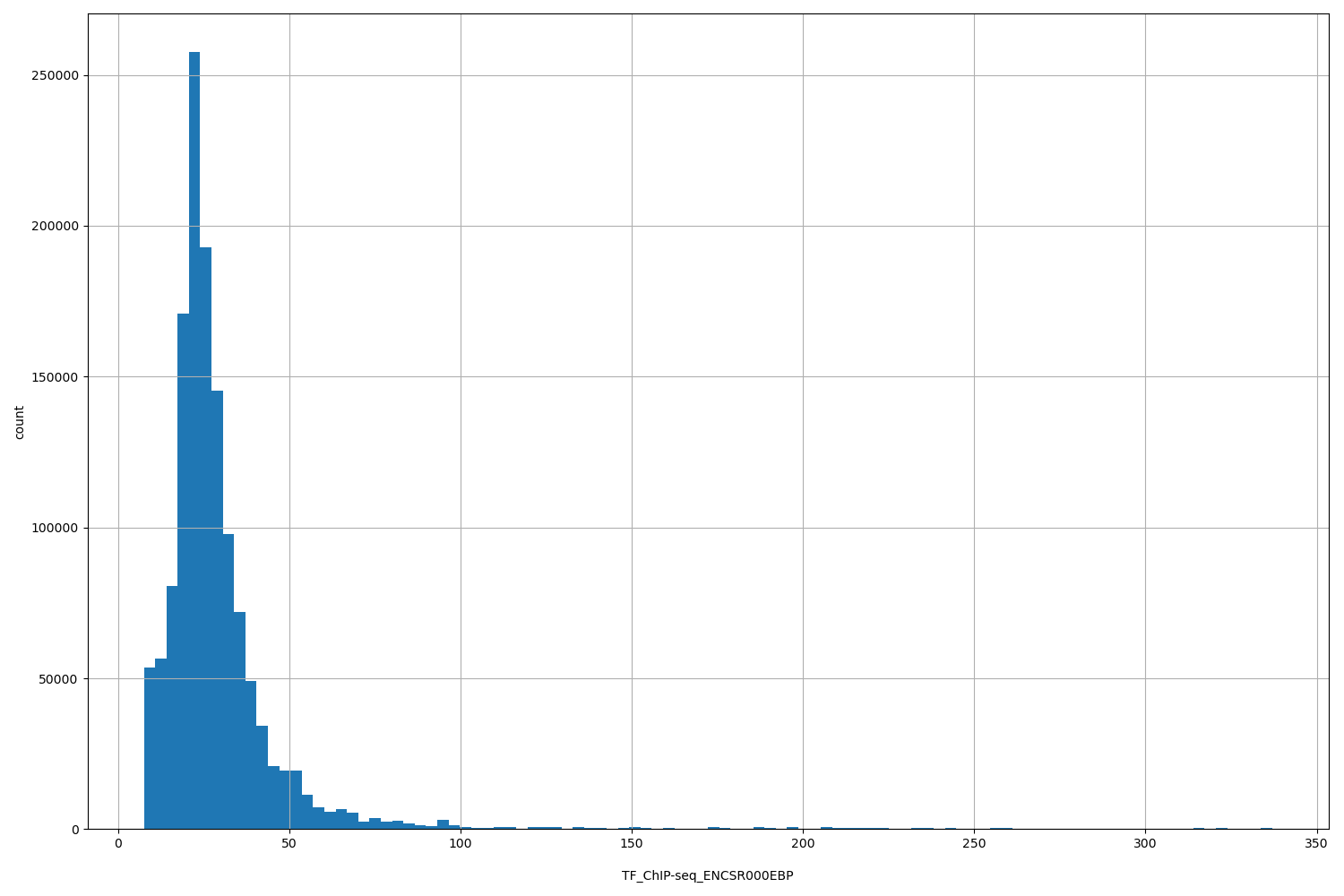
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EBP | float |
TF_ChIP-seq_ENCSR000EBP |
TF_ChIP-seq ENCSR000EBP [biosample_summary="Homo sapiens H1" and target="GTF2F1"]
|
 |
[7.57, 337] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF205YOD.bed.gz | 75.2 KB | c922c340580a815e7117f367dda72e7b |
| ENCFF205YOD.bed.gz.dvc | 99.0 B | ecbcd5a34d7954d807d33c7593e0c2e1 |
| ENCFF205YOD.tabix.bed.gz | 50.79 KB | b6ea93d0e937d44ffb149fac0e92cd87 |
| ENCFF205YOD.tabix.bed.gz.dvc | 105.0 B | 4bb646f081a2e51045c4c9a3f24610f3 |
| ENCFF205YOD.tabix.bed.gz.tbi | 38.62 KB | 27f4470400a44f4f72215029176046b6 |
| ENCFF205YOD.tabix.bed.gz.tbi.dvc | 109.0 B | 6f9cdb1f640bf6a5a5fd3c3913a25520 |
| genomic_resource.yaml | 2.96 KB | d5867519fa401a0b9a005248f956a7ea |
| genomic_resource_original.yaml | 2.87 KB | bb007887c0d2834ad8413444cf536d32 |
| statistics/ |