TF_ChIP-seq_ENCSR000EAJ
TF_ChIP-seq ENCSR000EAJ [biosample_summary="Homo sapiens GM12891" and target="POLR2A"]
| Id: | TF_ChIP-seq/ENCSR000EAJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EAJ [biosamplesummary="Homo sapiens GM12891" and target="POLR2A"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CQA|/files/ENCFF002CQA/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
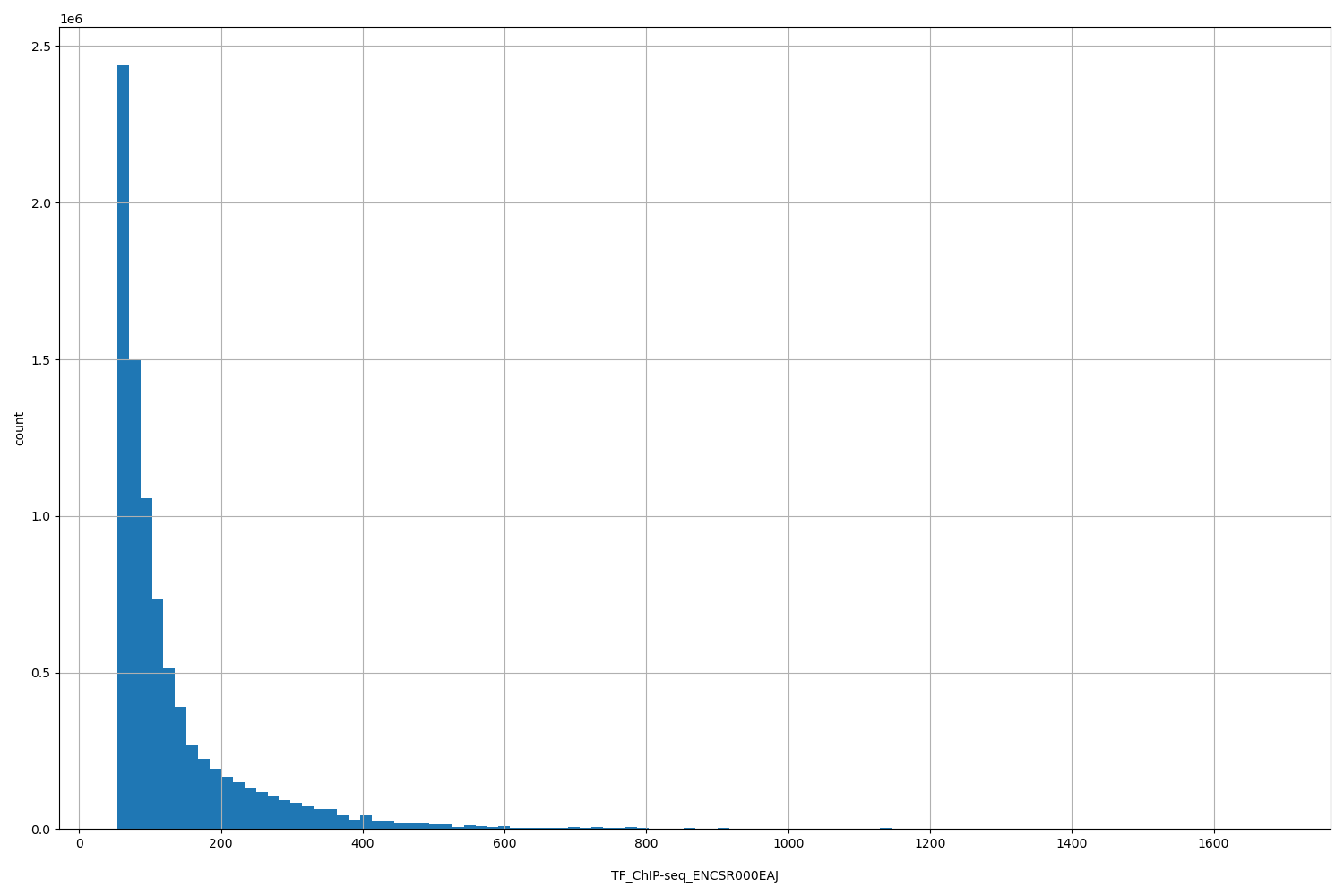
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EAJ | float |
TF_ChIP-seq_ENCSR000EAJ |
TF_ChIP-seq ENCSR000EAJ [biosample_summary="Homo sapiens GM12891" and target="POLR2A"]
|
 |
[53.9, 1.68e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CQA.bed.gz | 448.47 KB | 3e3edcca4591f6adc4da3f652692f702 |
| ENCFF002CQA.bed.gz.dvc | 100.0 B | a906e23876b335a042c0758e5d096217 |
| ENCFF002CQA.tabix.bed.gz | 342.83 KB | 639813a75a0bf16fe685544dbfef157c |
| ENCFF002CQA.tabix.bed.gz.dvc | 106.0 B | a600e618a8505fb6d6e67a36dc7df772 |
| ENCFF002CQA.tabix.bed.gz.tbi | 110.61 KB | 2f813f7077b13a325020a16e7a570d67 |
| ENCFF002CQA.tabix.bed.gz.tbi.dvc | 110.0 B | 7f8a267d9a1bf4817275f69441141871 |
| genomic_resource.yaml | 1.48 KB | 0a3da89e906272aef4c41f079a322123 |
| genomic_resource_original.yaml | 1.38 KB | 77b2128dc5f2aede6aa9771ddfc39afe |
| statistics/ |