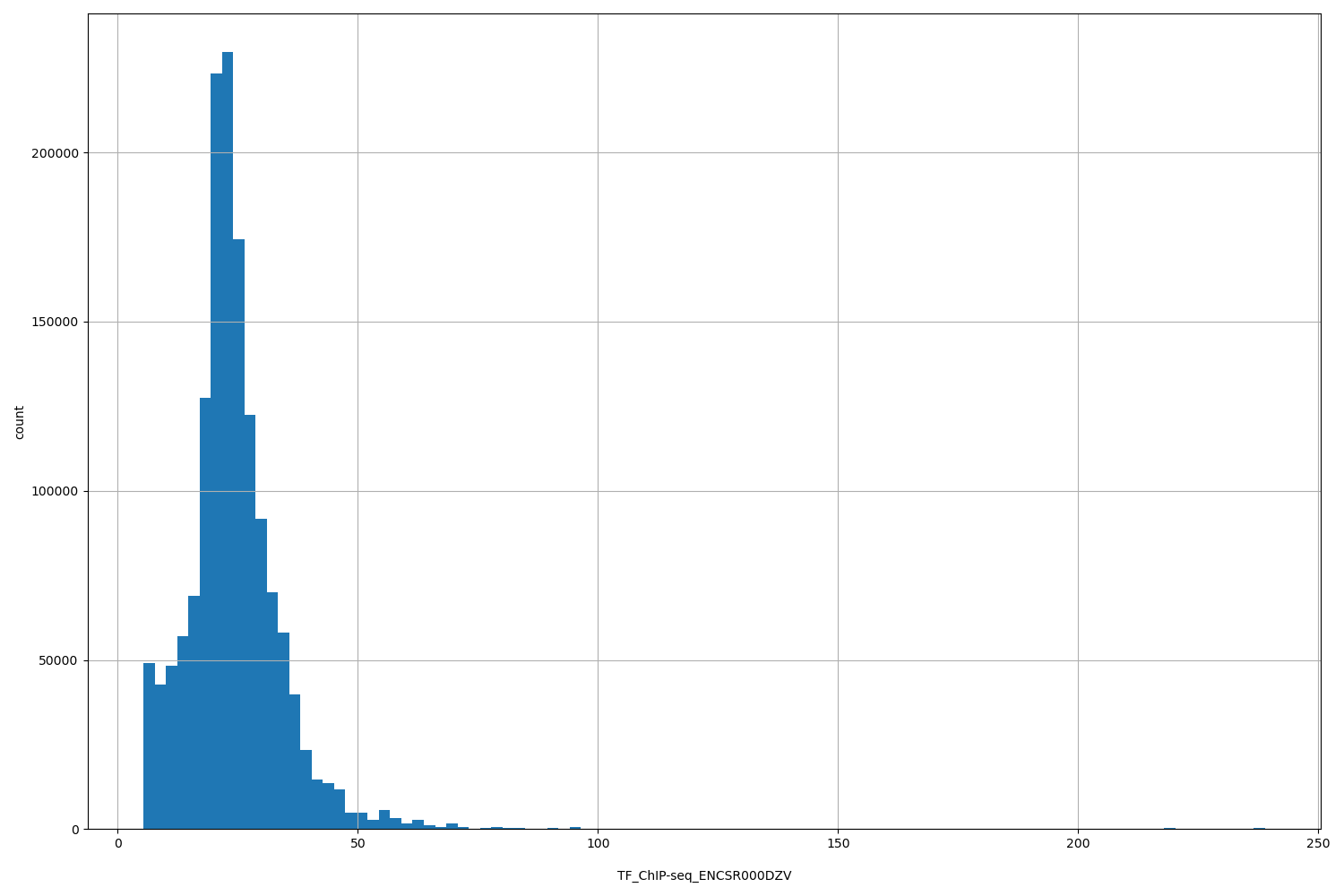
TF_ChIP-seq_ENCSR000DZV
| Id: | TF_ChIP-seq/ENCSR000DZV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DZV [biosamplesummary="Homo sapiens GM12878" and target="STAT3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN543ZZD|/analyses/ENCAN543ZZD/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN543ZZD|/analyses/ENCAN543ZZD/} has in progress subobject document {545c724b-f8cf-43d3-b326-9bd513bd1364|/documents/545c724b-f8cf-43d3-b326-9bd513bd1364/} audit_warning: Processed alignments file {ENCFF323QPE|/files/ENCFF323QPE/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16216683 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting STAT3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF226YHB|/files/ENCFF226YHB/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 15711742 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting STAT3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DZV | float |
TF_ChIP-seq_ENCSR000DZV |
TF_ChIP-seq ENCSR000DZV [biosample_summary="Homo sapiens GM12878" and target="STAT3"]
|
 |
[5.34, 239] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF029PAA.bed.gz | 75.41 KB | 6b8c2a2a3308bcf920a0aefd7379018b |
| ENCFF029PAA.bed.gz.dvc | 99.0 B | 77305437831b5d17666746e4ac93c5f9 |
| ENCFF029PAA.tabix.bed.gz | 56.03 KB | eb901cbf33e72d3dd06da8c6c45aa8bf |
| ENCFF029PAA.tabix.bed.gz.dvc | 105.0 B | e8075dabee408921bf19d68dfbb556a4 |
| ENCFF029PAA.tabix.bed.gz.tbi | 38.68 KB | 672443ee7c66ae4674e5bc7edbbdcbfe |
| ENCFF029PAA.tabix.bed.gz.tbi.dvc | 109.0 B | 8b0278593550f56180670363b7134f5f |
| genomic_resource.yaml | 2.57 KB | 51392ca51d9d768a48cd7c2ed61d6685 |
| genomic_resource_original.yaml | 2.47 KB | dac3dce472e50b657718904d43d0aa91 |
| statistics/ |