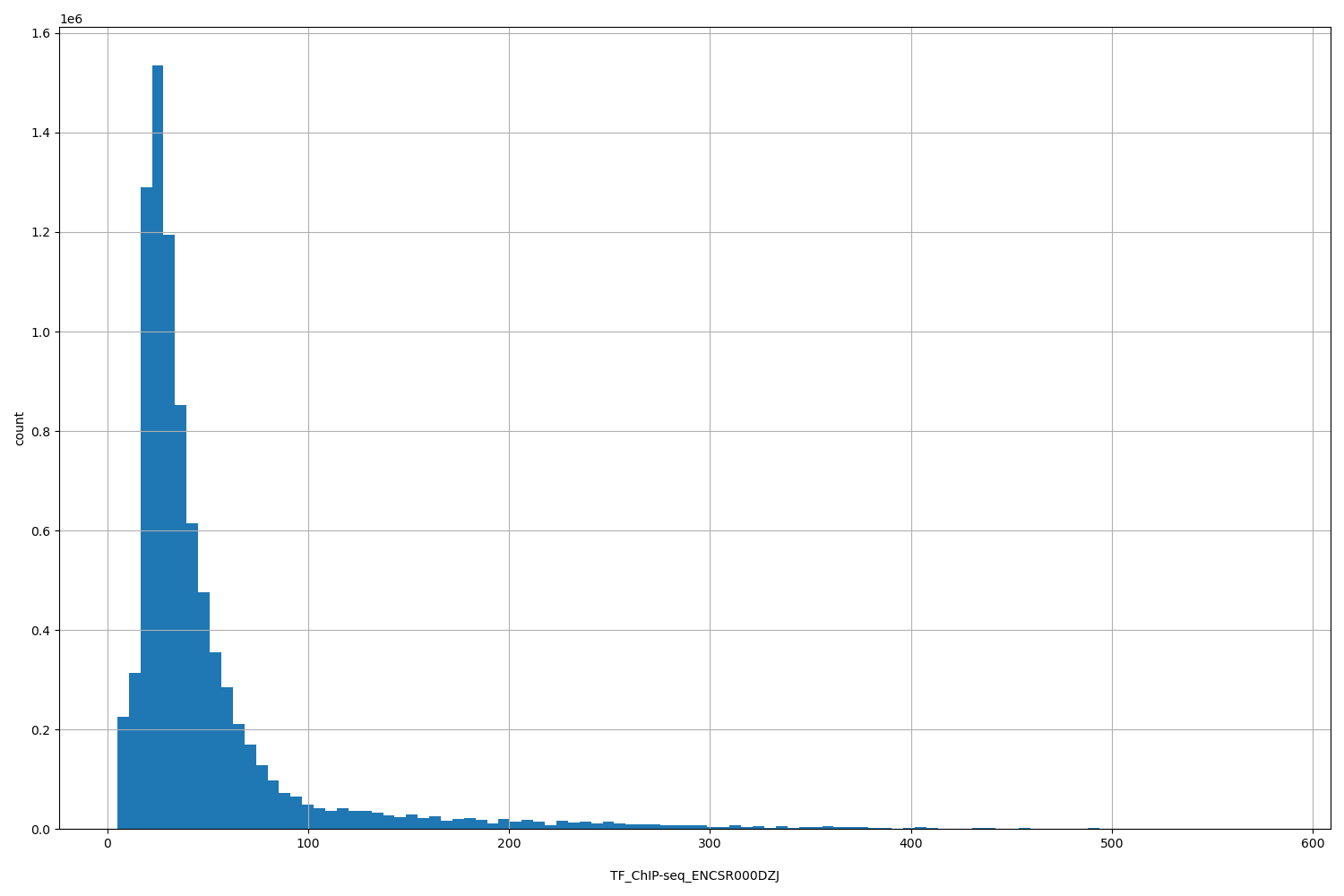
TF_ChIP-seq_ENCSR000DZJ
| Id: | TF_ChIP-seq/ENCSR000DZJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DZJ [biosamplesummary="Homo sapiens GM12878" and target="BHLHE40"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN943BBB|/analyses/ENCAN943BBB/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN943BBB|/analyses/ENCAN943BBB/} has in progress subobject document {fb8bcecf-ad07-4a9e-9ba1-9b14b38836b4|/documents/fb8bcecf-ad07-4a9e-9ba1-9b14b38836b4/} audit_warning: Processed alignments file {ENCFF947HBR|/files/ENCFF947HBR/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14578919 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BHLHE40-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF594ZIL|/files/ENCFF594ZIL/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18869238 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BHLHE40-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DZJ | float |
TF_ChIP-seq_ENCSR000DZJ |
TF_ChIP-seq ENCSR000DZJ [biosample_summary="Homo sapiens GM12878" and target="BHLHE40"]
|
 |
[5.05, 580] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF633YUQ.bed.gz | 490.52 KB | 83f8660a9ed7d3b129533a5e31b14d28 |
| ENCFF633YUQ.bed.gz.dvc | 100.0 B | 4a23681f7d022f8f985e3f59193e8102 |
| ENCFF633YUQ.tabix.bed.gz | 347.73 KB | 9ba0355e66def7e9fe1be780bcab07e9 |
| ENCFF633YUQ.tabix.bed.gz.dvc | 106.0 B | d67f0e4ae07e0bd1960357189f939d21 |
| ENCFF633YUQ.tabix.bed.gz.tbi | 164.96 KB | 964eed4c8abf48dc6b0c111a02c22342 |
| ENCFF633YUQ.tabix.bed.gz.tbi.dvc | 110.0 B | 648f00280ca77df0bf991b8454f9ce10 |
| genomic_resource.yaml | 2.58 KB | 659bf95efd2f6d1145f3c49df0af6b37 |
| genomic_resource_original.yaml | 2.48 KB | 2ee7fc6e106dff9931650e3400937066 |
| statistics/ |