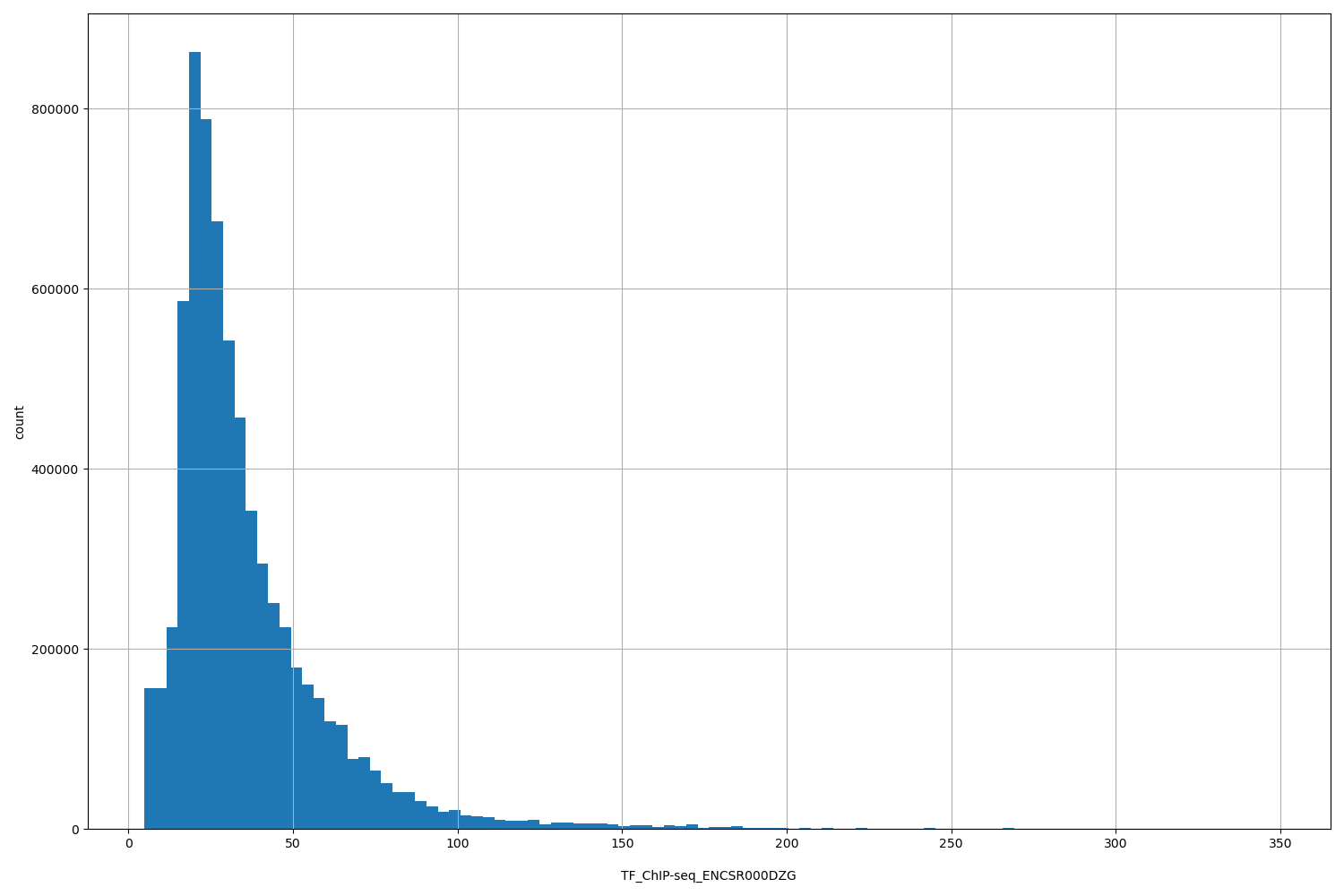
TF_ChIP-seq_ENCSR000DZG
| Id: | TF_ChIP-seq/ENCSR000DZG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DZG [biosamplesummary="Homo sapiens GM12878" and target="EP300"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN124EFK|/analyses/ENCAN124EFK/} has in progress subobject document {68e9615f-2b03-4942-b14a-d5668bcb9337|/documents/68e9615f-2b03-4942-b14a-d5668bcb9337/} audit_internal_action: Released analysis {ENCAN124EFK|/analyses/ENCAN124EFK/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF908KIL|/files/ENCFF908KIL/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14152656 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EP300-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF617MKT|/files/ENCFF617MKT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 15056566 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EP300-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF926AKK|/files/ENCFF926AKK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.22 and a self consistency ratio of 2.09. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DZG | float |
TF_ChIP-seq_ENCSR000DZG |
TF_ChIP-seq ENCSR000DZG [biosample_summary="Homo sapiens GM12878" and target="EP300"]
|
 |
[4.67, 348] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF926AKK.bed.gz | 349.97 KB | 84451db4ff5d07f36a72061f44800d33 |
| ENCFF926AKK.bed.gz.dvc | 100.0 B | cff433c4a658debdf475ed206cc8d30c |
| ENCFF926AKK.tabix.bed.gz | 259.65 KB | c4d46df30a653b5da222ec85fbe83b2d |
| ENCFF926AKK.tabix.bed.gz.dvc | 106.0 B | b8783224b606b4dcd51d3cb2cc7f35f6 |
| ENCFF926AKK.tabix.bed.gz.tbi | 131.82 KB | 15fc86f6d87cc6e5e84b19364b07ca0a |
| ENCFF926AKK.tabix.bed.gz.tbi.dvc | 110.0 B | 19912e3604cde14e66114b4dcda38a93 |
| genomic_resource.yaml | 3.11 KB | de5bbb8c8de353caeb46d70d747cc31b |
| genomic_resource_original.yaml | 3.01 KB | a8a1364dadbfd64ecd47c24a3e20a426 |
| statistics/ |