TF_ChIP-seq_ENCSR000DVA
| Id: | TF_ChIP-seq/ENCSR000DVA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DVA [biosamplesummary="Homo sapiens fibroblast of lung" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF777ODE|/files/ENCFF777ODE/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN275TFY|/analyses/ENCAN275TFY/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF327QNZ|/files/ENCFF327QNZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 17549347 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
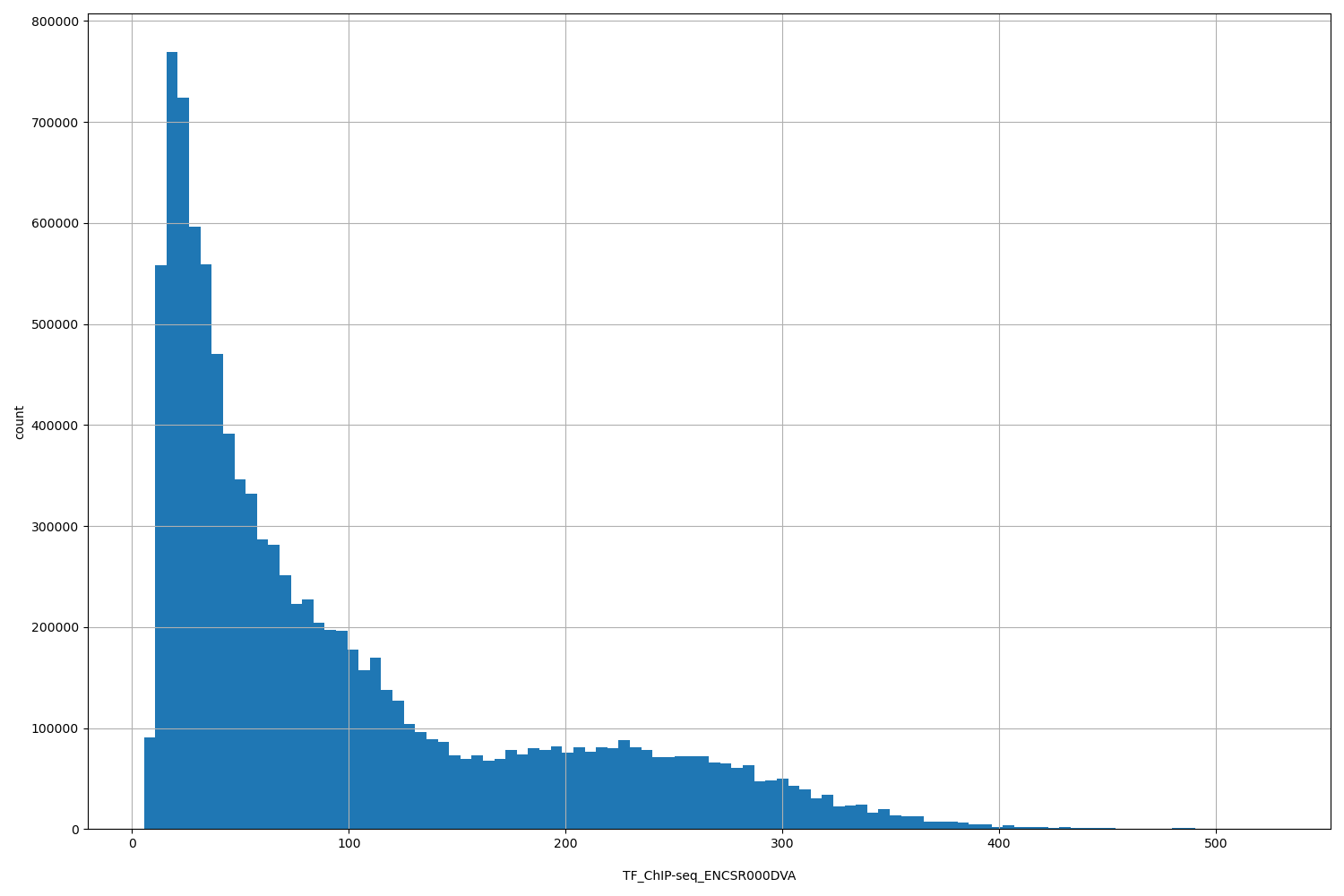
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DVA | float |
TF_ChIP-seq_ENCSR000DVA |
TF_ChIP-seq ENCSR000DVA [biosample_summary="Homo sapiens fibroblast of lung" and target="CTCF"]
|
 |
[5.43, 527] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF777ODE.bed.gz | 746.15 KB | 325603dfc60c0749af80c8a6366daad9 |
| ENCFF777ODE.bed.gz.dvc | 100.0 B | fce11a0e5bdc91c308355659ce6126a3 |
| ENCFF777ODE.tabix.bed.gz | 543.68 KB | 97a35fa76c29bfd8e2d99da9ef3886bd |
| ENCFF777ODE.tabix.bed.gz.dvc | 106.0 B | 0bc640a22df7de58069dfddf108fd29a |
| ENCFF777ODE.tabix.bed.gz.tbi | 286.44 KB | d790d924201ec630169ba210bb94db9d |
| ENCFF777ODE.tabix.bed.gz.tbi.dvc | 110.0 B | 7fa5528b4b9c6dbfef1cf198241eb02e |
| genomic_resource.yaml | 2.06 KB | d12056e30c4dd61886aeaaa84e6a0710 |
| genomic_resource_original.yaml | 1.96 KB | a31562f271ada566594231843a0d745a |
| statistics/ |