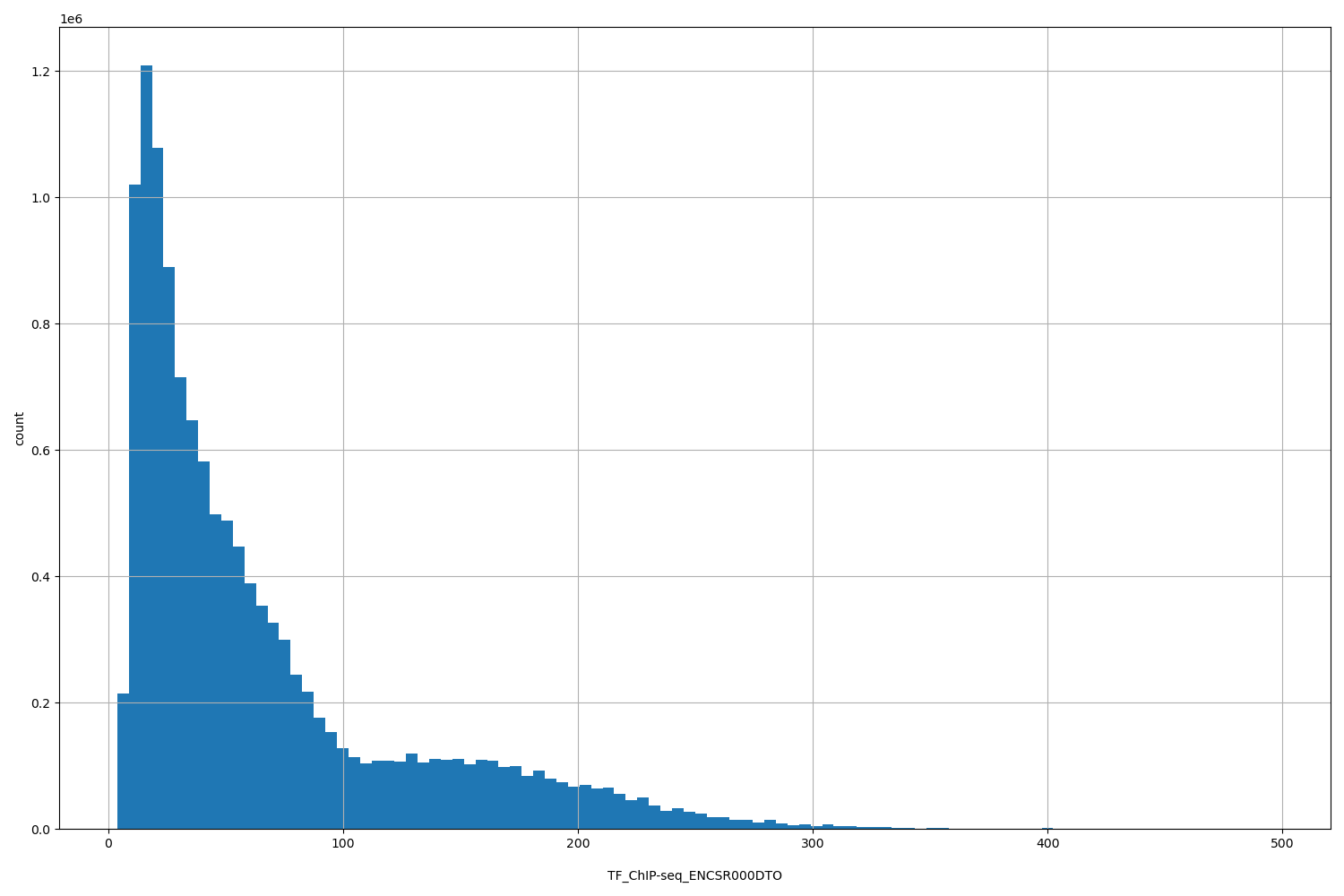
TF_ChIP-seq_ENCSR000DTO
| Id: | TF_ChIP-seq/ENCSR000DTO |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DTO [biosamplesummary="Homo sapiens HCT116" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF171SNH|/files/ENCFF171SNH/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN334FCK|/analyses/ENCAN334FCK/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF091ODJ|/files/ENCFF091ODJ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 13580424 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF648CAE|/files/ENCFF648CAE/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 11606264 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CTCF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DTO | float |
TF_ChIP-seq_ENCSR000DTO |
TF_ChIP-seq ENCSR000DTO [biosample_summary="Homo sapiens HCT116" and target="CTCF"]
|
 |
[3.89, 496] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF171SNH.bed.gz | 793.74 KB | 7c30b0282011df4c684ff487beab76bc |
| ENCFF171SNH.bed.gz.dvc | 100.0 B | 8f9fbf6c90b910662b0bf5340335ee58 |
| ENCFF171SNH.tabix.bed.gz | 613.96 KB | 02d1718d47dba3bae0e3547c6088e58e |
| ENCFF171SNH.tabix.bed.gz.dvc | 106.0 B | 2052838d43869a29cc9aa8446fd9d129 |
| ENCFF171SNH.tabix.bed.gz.tbi | 316.58 KB | 1bbf28f9e3933ae2b8a00d61aea12ec2 |
| ENCFF171SNH.tabix.bed.gz.tbi.dvc | 110.0 B | 2c44b8cb6ea211f68735d3d3d34d7fa8 |
| genomic_resource.yaml | 2.72 KB | 11f76ea53c0df9eb812d0171f4cfaedf |
| genomic_resource_original.yaml | 2.63 KB | a92d621bc20521fff4ddd38f42d08325 |
| statistics/ |