TF_ChIP-seq_ENCSR000DOH
| Id: | TF_ChIP-seq/ENCSR000DOH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DOH [biosamplesummary="Homo sapiens K562" and target="SIRT6"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN243JIY|/analyses/ENCAN243JIY/} has in progress subobject document {b1efc2f0-7ff4-4186-91a3-f11c14162d7e|/documents/b1efc2f0-7ff4-4186-91a3-f11c14162d7e/} audit_internal_action: Released analysis {ENCAN243JIY|/analyses/ENCAN243JIY/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF321WQD|/files/ENCFF321WQD/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6178753 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SIRT6-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF645ZIX|/files/ENCFF645ZIX/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6038193 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SIRT6-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
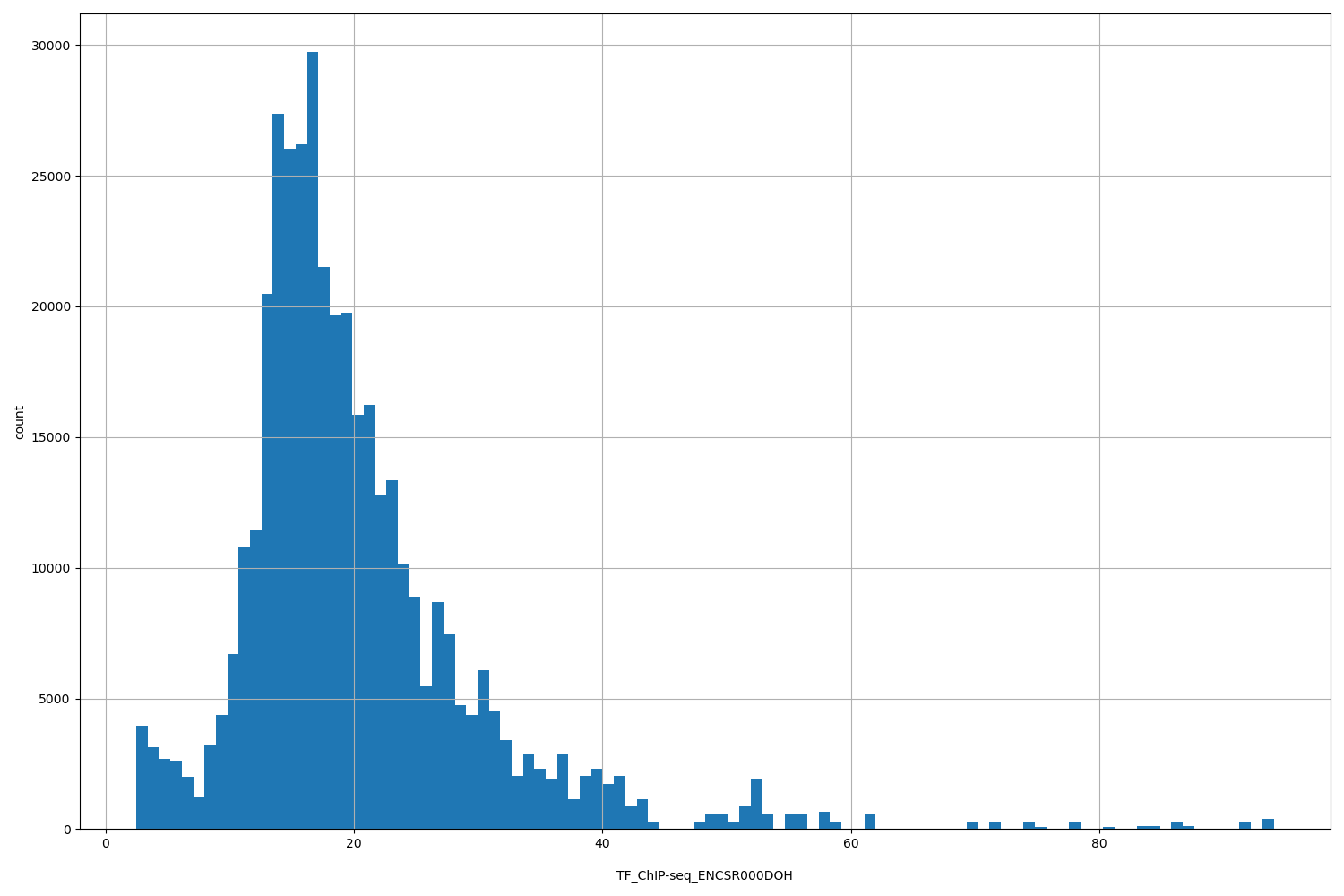
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DOH | float |
TF_ChIP-seq_ENCSR000DOH |
TF_ChIP-seq ENCSR000DOH [biosample_summary="Homo sapiens K562" and target="SIRT6"]
|
 |
[2.47, 94] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF217ACS.bed.gz | 24.56 KB | f195c93166b6bbaca39773eb191957c7 |
| ENCFF217ACS.bed.gz.dvc | 99.0 B | 48d983d7db92e9da0c889904b9c591d2 |
| ENCFF217ACS.tabix.bed.gz | 17.46 KB | 83a6bd129ffa84432e29152f3335543a |
| ENCFF217ACS.tabix.bed.gz.dvc | 105.0 B | ac63bfa48a62d9e690618ea33f10b218 |
| ENCFF217ACS.tabix.bed.gz.tbi | 18.23 KB | e4efd14143a98c319312769d98e70260 |
| ENCFF217ACS.tabix.bed.gz.tbi.dvc | 109.0 B | 581b839d8bcadf574220703313bfde84 |
| genomic_resource.yaml | 2.57 KB | 5ead65fb707b6dd5db213f3a97a8351d |
| genomic_resource_original.yaml | 2.48 KB | 8a3632f9f6c046157d0b0e25b270ae8c |
| statistics/ |