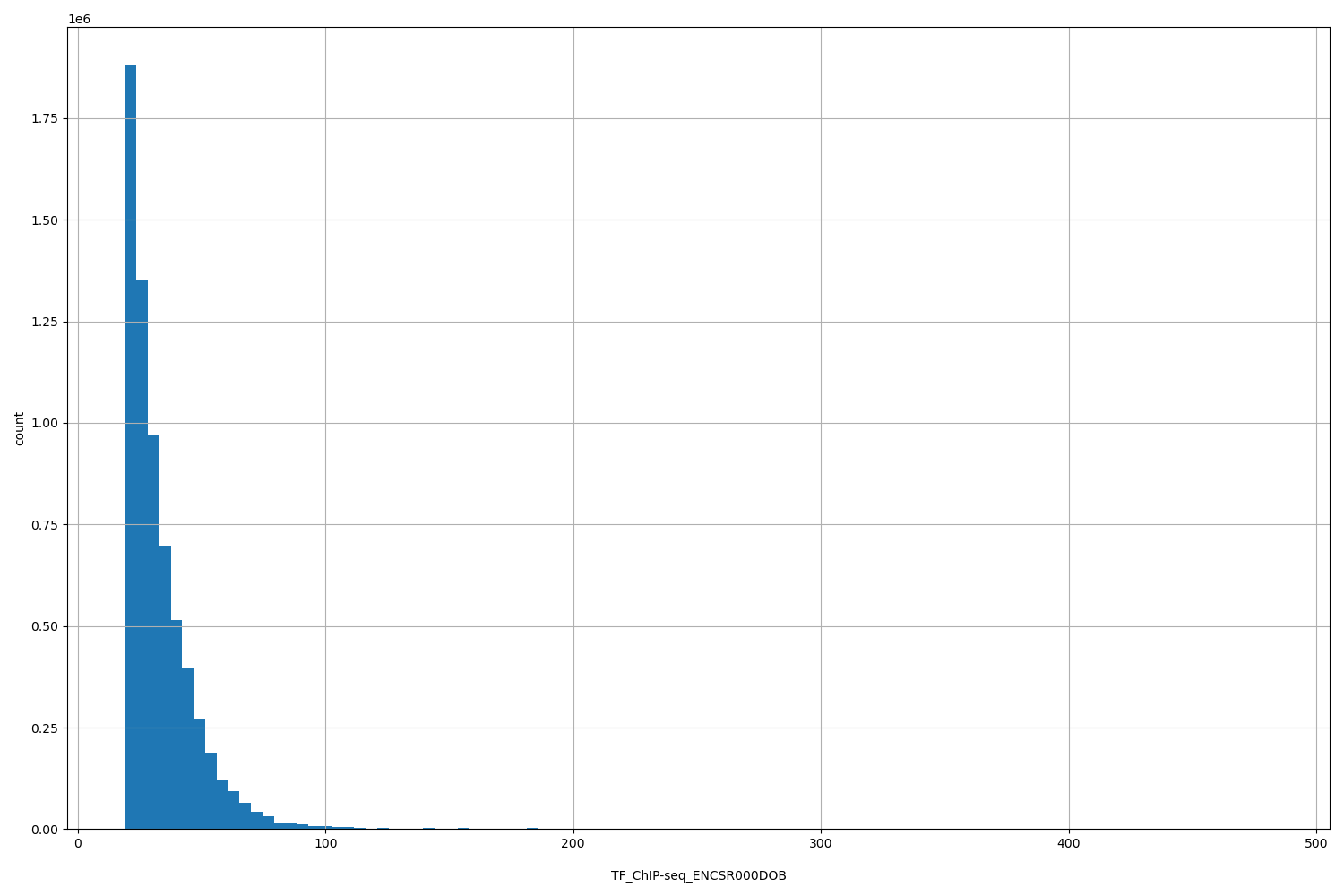
TF_ChIP-seq_ENCSR000DOB
TF_ChIP-seq ENCSR000DOB [biosample_summary="Homo sapiens K562" and target="HMGN3"]
| Id: | TF_ChIP-seq/ENCSR000DOB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DOB [biosamplesummary="Homo sapiens K562" and target="HMGN3"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CWT|/files/ENCFF002CWT/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DOB | float |
TF_ChIP-seq_ENCSR000DOB |
TF_ChIP-seq ENCSR000DOB [biosample_summary="Homo sapiens K562" and target="HMGN3"]
|
 |
[18.9, 482] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CWT.bed.gz | 313.22 KB | 9784935821cde1c75e773aba4c415e7d |
| ENCFF002CWT.bed.gz.dvc | 100.0 B | 5dd6a15a330252c7df414263610c7a90 |
| ENCFF002CWT.tabix.bed.gz | 244.9 KB | 9d167529bd520dce9c6961580a6b66d2 |
| ENCFF002CWT.tabix.bed.gz.dvc | 106.0 B | 2eefcf4e74a93430ac1f3bd49cdedf24 |
| ENCFF002CWT.tabix.bed.gz.tbi | 98.62 KB | 0b5d862a32462c052a2a37c984d1403b |
| ENCFF002CWT.tabix.bed.gz.tbi.dvc | 110.0 B | bd965370f2594b3dedd41b0b6b494c74 |
| genomic_resource.yaml | 1.47 KB | fad02a6b8d258853215012b6d66f354c |
| genomic_resource_original.yaml | 1.38 KB | 18e12e53549743306f01a04baaa5baab |
| statistics/ |