TF_ChIP-seq_ENCSR000DNX
TF_ChIP-seq ENCSR000DNX [biosample_summary="Homo sapiens HeLa-S3" and target="BDP1"]
| Id: | TF_ChIP-seq/ENCSR000DNX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DNX [biosamplesummary="Homo sapiens HeLa-S3" and target="BDP1"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CRV|/files/ENCFF002CRV/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
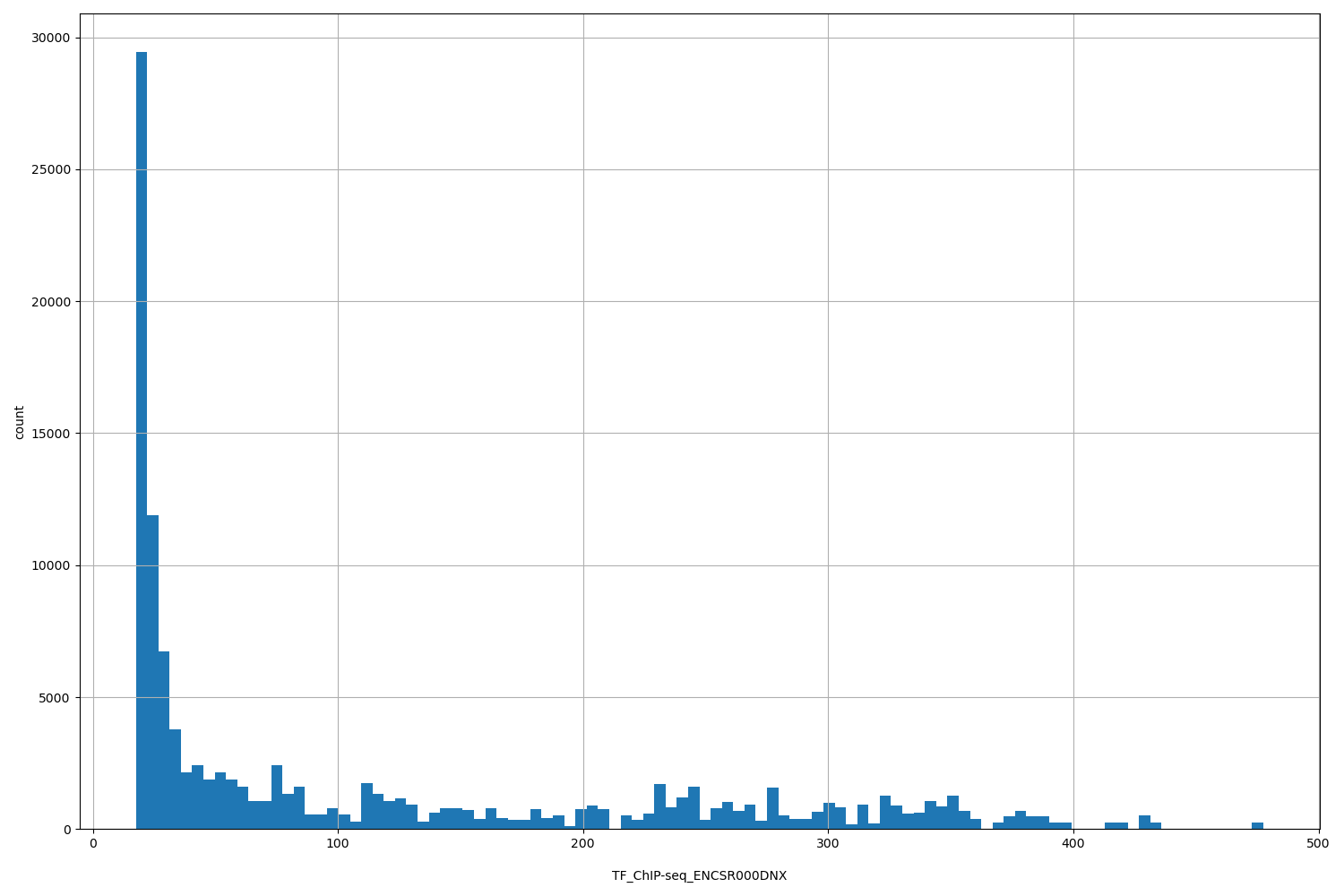
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DNX | float |
TF_ChIP-seq_ENCSR000DNX |
TF_ChIP-seq ENCSR000DNX [biosample_summary="Homo sapiens HeLa-S3" and target="BDP1"]
|
 |
[17.6, 477] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CRV.bed.gz | 12.92 KB | fe274615f1addff9d92690272a00fbc4 |
| ENCFF002CRV.bed.gz.dvc | 99.0 B | f92fb7cec120034b2453215c35167cde |
| ENCFF002CRV.tabix.bed.gz | 7.94 KB | 4ac7f1c7e57d3e3d70d49e0c0257b2c1 |
| ENCFF002CRV.tabix.bed.gz.dvc | 104.0 B | 801f4e7967fca1b6d7b850da9597bb1f |
| ENCFF002CRV.tabix.bed.gz.tbi | 8.53 KB | 6e09e951737788868683eef1df9608ea |
| ENCFF002CRV.tabix.bed.gz.tbi.dvc | 108.0 B | 4e18616520639873b8cb4fc576e9f71b |
| genomic_resource.yaml | 1.48 KB | 578f00eef13c9452913d17c860e9a83c |
| genomic_resource_original.yaml | 1.38 KB | db404e22bc1dda0d1bc14849a6415775 |
| statistics/ |