TF_ChIP-seq_ENCSR000DNT
| Id: | TF_ChIP-seq/ENCSR000DNT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DNT [biosamplesummary="Homo sapiens HeLa-S3" and target="ZZZ3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN694MIK|/analyses/ENCAN694MIK/} has in progress subobject document {0d288020-0f3d-48a2-baad-eda7e8e6a951|/documents/0d288020-0f3d-48a2-baad-eda7e8e6a951/} audit_internal_action: Released analysis {ENCAN694MIK|/analyses/ENCAN694MIK/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF719MUI|/files/ENCFF719MUI/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14436538 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZZZ3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF374OBP|/files/ENCFF374OBP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13984737 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZZZ3-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
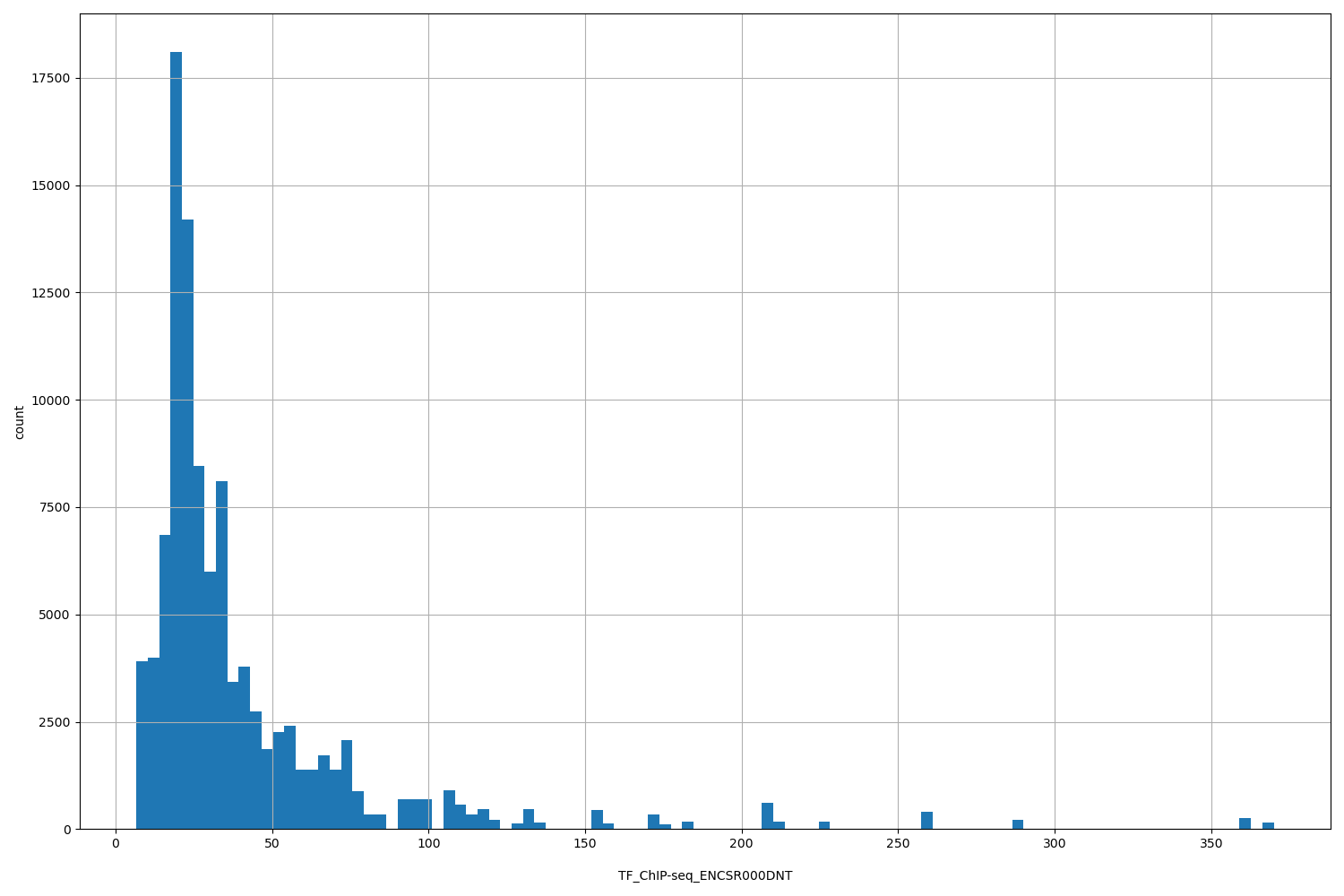
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DNT | float |
TF_ChIP-seq_ENCSR000DNT |
TF_ChIP-seq ENCSR000DNT [biosample_summary="Homo sapiens HeLa-S3" and target="ZZZ3"]
|
 |
[6.67, 370] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF675ARG.bed.gz | 6.4 KB | 07f309087dce974b8ea10d2f06439ac7 |
| ENCFF675ARG.bed.gz.dvc | 98.0 B | 7bf3ee6bc5261f004afe40df89f5fa0d |
| ENCFF675ARG.tabix.bed.gz | 4.17 KB | 5786052d42c34d6ad8fe1f4c9b67b212 |
| ENCFF675ARG.tabix.bed.gz.dvc | 104.0 B | 1751e56a1b21266b88a73d8aa690e127 |
| ENCFF675ARG.tabix.bed.gz.tbi | 7.4 KB | efee8636117aebd94eed288169d87ea3 |
| ENCFF675ARG.tabix.bed.gz.tbi.dvc | 108.0 B | d084429e4369e04d1ef96e99e89dde13 |
| genomic_resource.yaml | 2.56 KB | c56322392ead2880356381e7385456d8 |
| genomic_resource_original.yaml | 2.47 KB | 083a504b49baa45b9bdb406796e6721b |
| statistics/ |