TF_ChIP-seq_ENCSR000DMZ
TF_ChIP-seq ENCSR000DMZ [biosample_summary="Homo sapiens A549" and target="POLR2A"]
| Id: | TF_ChIP-seq/ENCSR000DMZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DMZ [biosamplesummary="Homo sapiens A549" and target="POLR2A"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002DAE|/files/ENCFF002DAE/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
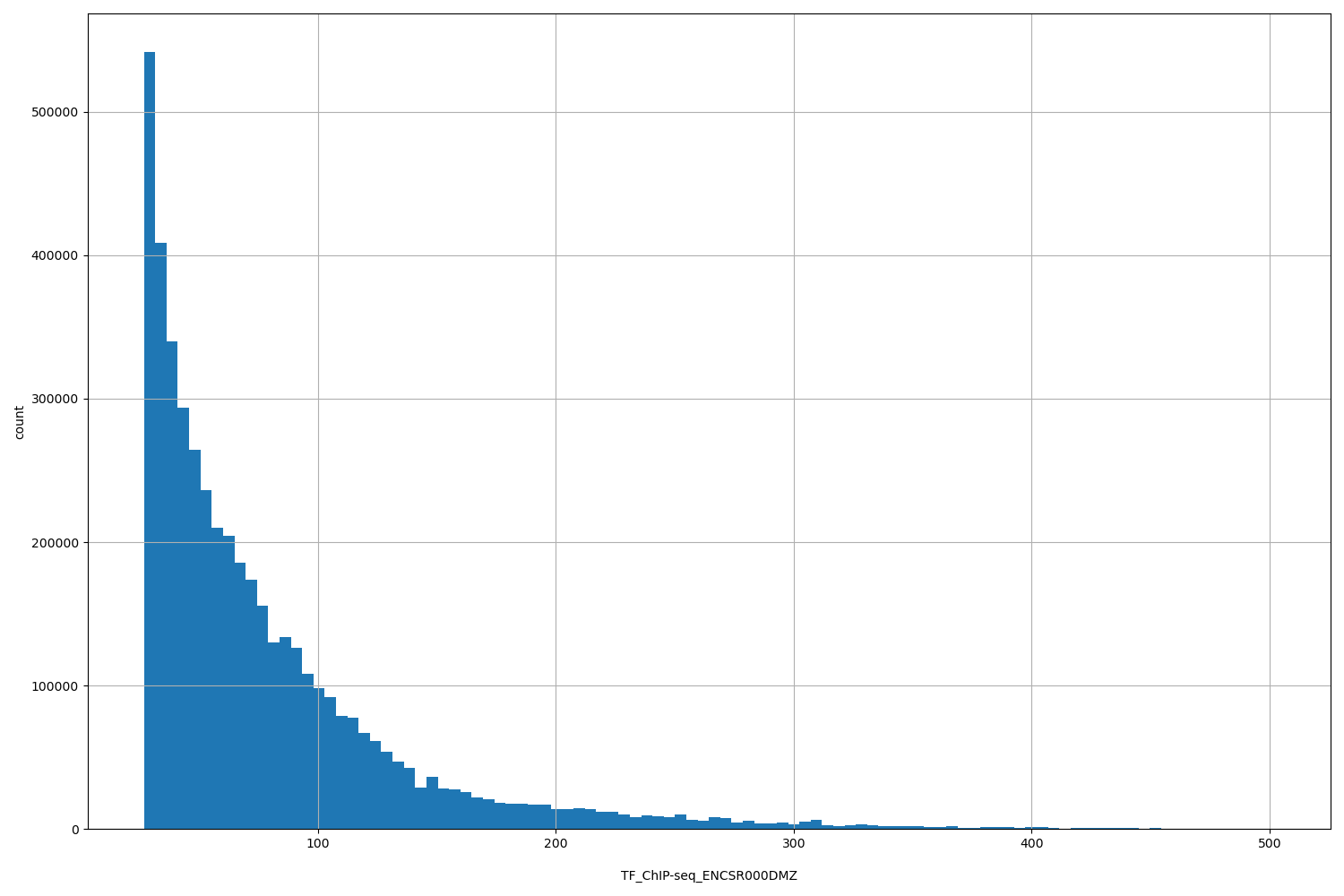
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DMZ | float |
TF_ChIP-seq_ENCSR000DMZ |
TF_ChIP-seq ENCSR000DMZ [biosample_summary="Homo sapiens A549" and target="POLR2A"]
|
 |
[27, 502] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002DAE.bed.gz | 353.92 KB | 9cb690330aa44a71ed0e54db45920aba |
| ENCFF002DAE.bed.gz.dvc | 100.0 B | 782f42ed6b83cd94902d0c8e98062474 |
| ENCFF002DAE.tabix.bed.gz | 268.2 KB | 3ac9457c4c2fff34ee50adcae365dadd |
| ENCFF002DAE.tabix.bed.gz.dvc | 106.0 B | 3e032ee1e58df96ae677a2e24a0c96ba |
| ENCFF002DAE.tabix.bed.gz.tbi | 106.93 KB | b666a5d3ccfa3cc0e8edd53a45205bd7 |
| ENCFF002DAE.tabix.bed.gz.tbi.dvc | 110.0 B | 43da942fc81731649e0e09c01fc661e5 |
| genomic_resource.yaml | 1.47 KB | cf2b900eb793445e85bbbfded2e8d382 |
| genomic_resource_original.yaml | 1.38 KB | 02732ab30ac7b14f9856d1bf4f3740cd |
| statistics/ |