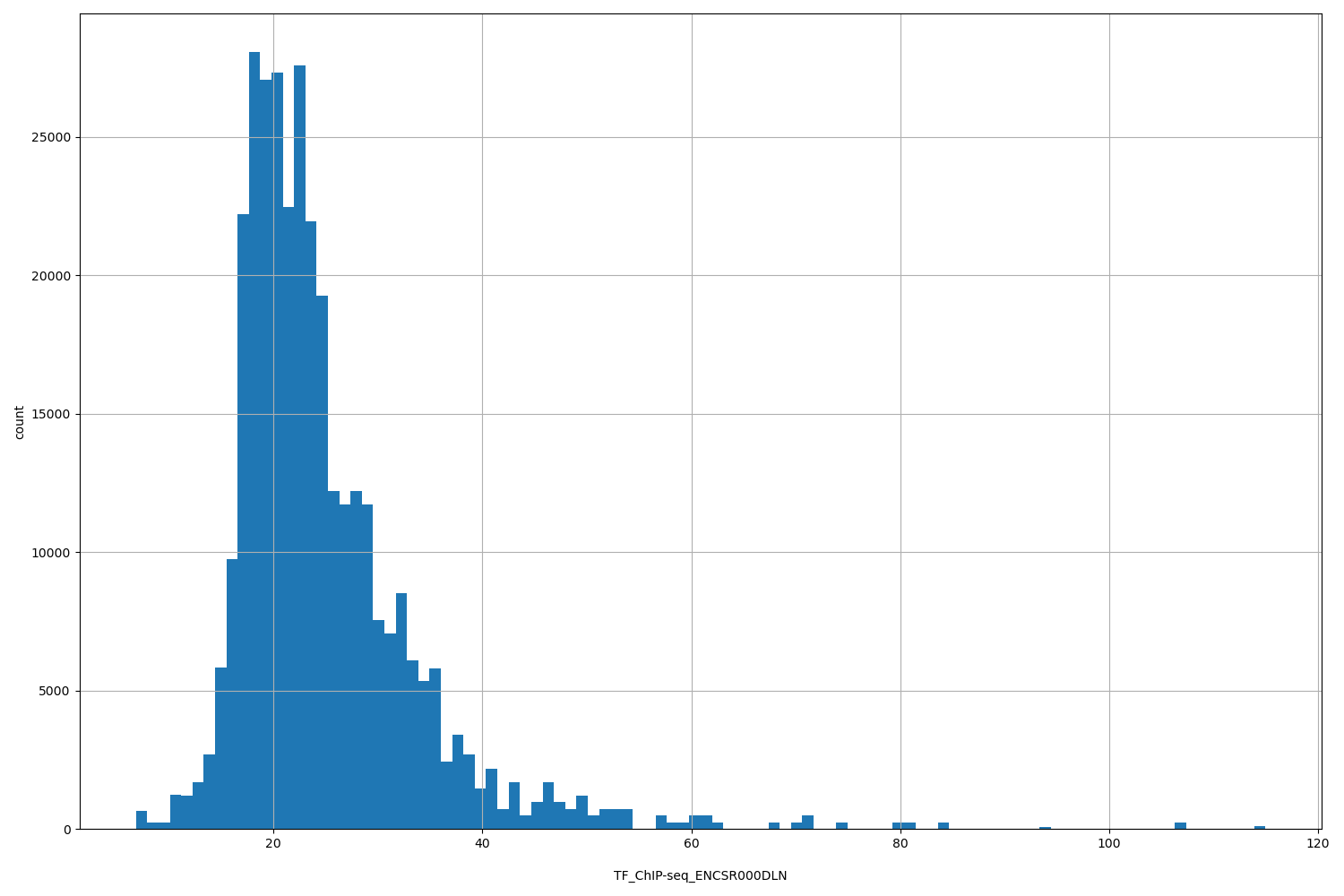
TF_ChIP-seq_ENCSR000DLN
| Id: | TF_ChIP-seq/ENCSR000DLN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000DLN [biosamplesummary="Homo sapiens HeLa-S3" and target="MYC"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN564AAU|/analyses/ENCAN564AAU/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN564AAU|/analyses/ENCAN564AAU/} has in progress subobject document {f09c2722-325c-411b-9303-8d47affedc31|/documents/f09c2722-325c-411b-9303-8d47affedc31/} audit_not_compliant: Processed alignments file {ENCFF237MZO|/files/ENCFF237MZO/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 5235893 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF620MEQ|/files/ENCFF620MEQ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 5374464 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MYC-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF784BWK|/files/ENCFF784BWK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.06 and a self consistency ratio of 2.61. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000DLN | float |
TF_ChIP-seq_ENCSR000DLN |
TF_ChIP-seq ENCSR000DLN [biosample_summary="Homo sapiens HeLa-S3" and target="MYC"]
|
 |
[6.83, 115] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF784BWK.bed.gz | 22.83 KB | cc3efc91634aba6631367f2dba20146f |
| ENCFF784BWK.bed.gz.dvc | 99.0 B | 9dd77c05bfaf2281d582666b3c614adf |
| ENCFF784BWK.tabix.bed.gz | 16.67 KB | d7eec21ee290d26520599ec199bf3931 |
| ENCFF784BWK.tabix.bed.gz.dvc | 105.0 B | 3853b6185bd4a974ff8abd1dc5c06094 |
| ENCFF784BWK.tabix.bed.gz.tbi | 18.72 KB | 467a213a91e5de12003d4808d56c1e16 |
| ENCFF784BWK.tabix.bed.gz.tbi.dvc | 109.0 B | 6105bbb8e4410b7a0ea160530770e350 |
| genomic_resource.yaml | 3.12 KB | f1933fb482875a31a72b0dc15edc36a6 |
| genomic_resource_original.yaml | 3.02 KB | 7953313da378576d7c0703b22267e39e |
| statistics/ |