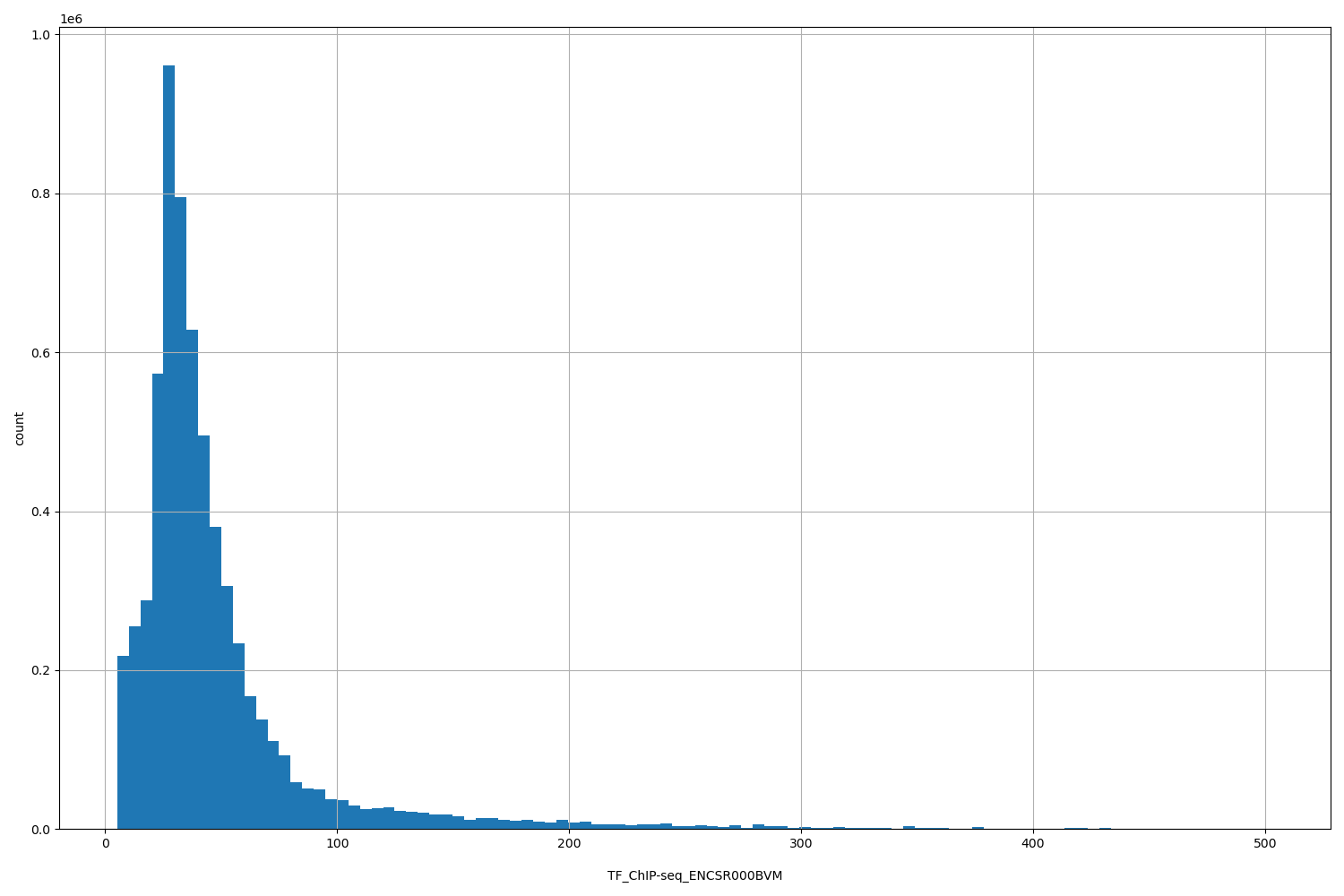
TF_ChIP-seq_ENCSR000BVM
| Id: | TF_ChIP-seq/ENCSR000BVM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BVM [biosamplesummary="Homo sapiens HepG2" and target="NR2F2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN613XZV|/analyses/ENCAN613XZV/} has in progress subobject document {9373d5b1-514c-4d41-8f9c-5f68e9593b66|/documents/9373d5b1-514c-4d41-8f9c-5f68e9593b66/} audit_internal_action: Released analysis {ENCAN613XZV|/analyses/ENCAN613XZV/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF485EPH|/files/ENCFF485EPH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 12655855 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NR2F2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF804MKP|/files/ENCFF804MKP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.42 and a self consistency ratio of 2.55. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BVM | float |
TF_ChIP-seq_ENCSR000BVM |
TF_ChIP-seq ENCSR000BVM [biosample_summary="Homo sapiens HepG2" and target="NR2F2"]
|
 |
[5.18, 503] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF804MKP.bed.gz | 320.68 KB | d47f4a861c40fab8d31ce136bc707512 |
| ENCFF804MKP.bed.gz.dvc | 100.0 B | 3b44bcd6bbf865369d00eb5044f472e6 |
| ENCFF804MKP.tabix.bed.gz | 231.11 KB | 01f82bbb0cf0e936439bca71f1a3bc84 |
| ENCFF804MKP.tabix.bed.gz.dvc | 106.0 B | 4d57c90a8b4ef051fbe94ab6c67c8ed1 |
| ENCFF804MKP.tabix.bed.gz.tbi | 115.09 KB | ef1d39df80576f8917f39a0d8bda1c90 |
| ENCFF804MKP.tabix.bed.gz.tbi.dvc | 110.0 B | 027eabf270a9a0ed6cc2210407e7ddb5 |
| genomic_resource.yaml | 2.6 KB | 6fd0640ee6cf3adc7c0e0aeb13d28418 |
| genomic_resource_original.yaml | 2.51 KB | d67cd2fc8c76c0c1200893b89739b3c7 |
| statistics/ |