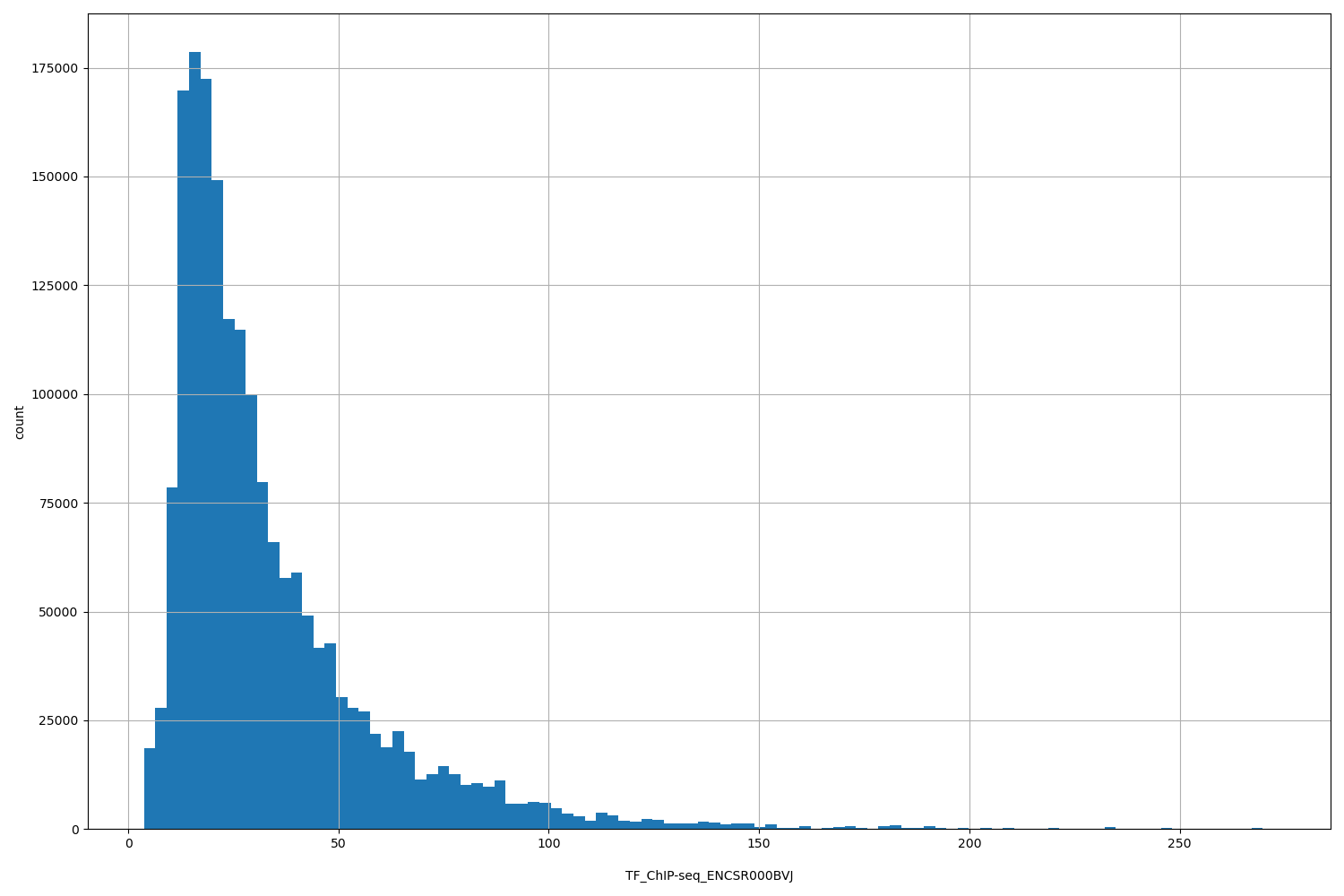
TF_ChIP-seq_ENCSR000BVJ
| Id: | TF_ChIP-seq/ENCSR000BVJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BVJ [biosamplesummary="Homo sapiens HCT116" and target="TEAD4"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN214VZS|/analyses/ENCAN214VZS/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN214VZS|/analyses/ENCAN214VZS/} has in progress subobject document {a5d95c70-7c97-4fc1-aed4-45f284c15a7d|/documents/a5d95c70-7c97-4fc1-aed4-45f284c15a7d/} audit_warning: Processed alignments file {ENCFF520EON|/files/ENCFF520EON/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 16587487 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting TEAD4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF391WGH|/files/ENCFF391WGH/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 11982749 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting TEAD4-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BVJ | float |
TF_ChIP-seq_ENCSR000BVJ |
TF_ChIP-seq ENCSR000BVJ [biosample_summary="Homo sapiens HCT116" and target="TEAD4"]
|
 |
[3.72, 272] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF008LFE.bed.gz | 121.42 KB | 4600791aa1f5b9008a1f88fca7122632 |
| ENCFF008LFE.bed.gz.dvc | 100.0 B | 8ded20aad7b4e2d42791daae0e588576 |
| ENCFF008LFE.tabix.bed.gz | 82.53 KB | bd24fa504fe11508aefcd6c404859113 |
| ENCFF008LFE.tabix.bed.gz.dvc | 105.0 B | 616b87e77c08d19b4ce54b0116a90613 |
| ENCFF008LFE.tabix.bed.gz.tbi | 62.05 KB | 45d61b95d6d0420dc18181c15c2575ec |
| ENCFF008LFE.tabix.bed.gz.tbi.dvc | 109.0 B | 1f6f08a3e4288f949ff79bdd8c6be3bc |
| genomic_resource.yaml | 2.56 KB | 30cfa9de1b6661b886a5592b126cb5b0 |
| genomic_resource_original.yaml | 2.47 KB | ae99116fc91ffc498eaaf40022bd9cdd |
| statistics/ |