TF_ChIP-seq_ENCSR000BUE
| Id: | TF_ChIP-seq/ENCSR000BUE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BUE [biosamplesummary="Homo sapiens Ishikawa" and target="EP300"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN672VSD|/analyses/ENCAN672VSD/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF621FAY|/files/ENCFF621FAY/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19591800 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EP300-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF712OLL|/files/ENCFF712OLL/}, {ENCFF442TIY|/files/ENCFF442TIY/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.54 and a self consistency ratio of 3.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF765PON|/files/ENCFF765PON/}, {ENCFF506UZB|/files/ENCFF506UZB/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.54 and a self consistency ratio of 3.04. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
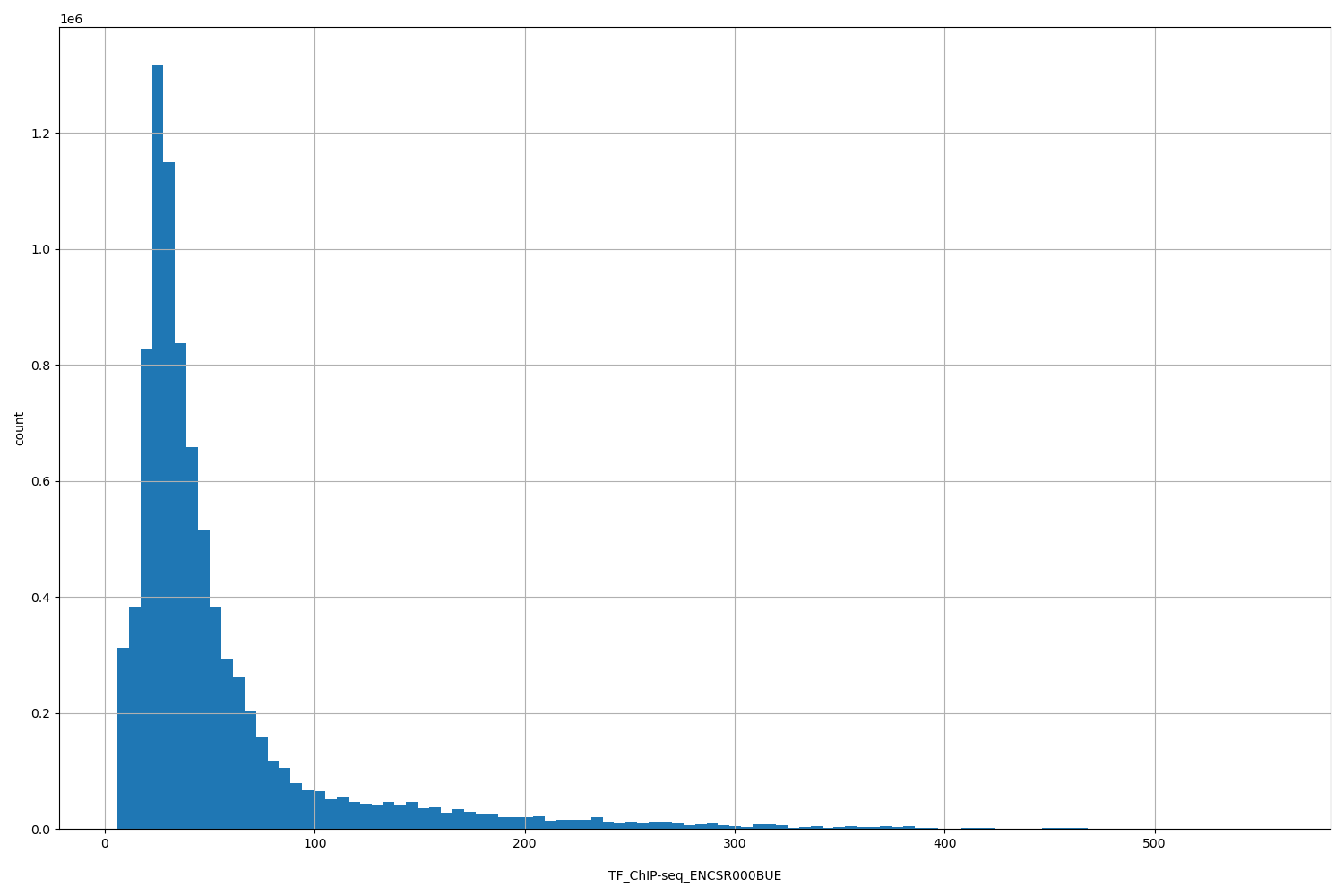
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BUE | float |
TF_ChIP-seq_ENCSR000BUE |
TF_ChIP-seq ENCSR000BUE [biosample_summary="Homo sapiens Ishikawa" and target="EP300"]
|
 |
[5.94, 556] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF765PON.bed.gz | 429.28 KB | 03643734a7b2745e034c1cfaf05a6863 |
| ENCFF765PON.bed.gz.dvc | 100.0 B | b2e0a98e9751cccfb426b7207ea6b236 |
| ENCFF765PON.tabix.bed.gz | 300.58 KB | 4445ce14551310642fcd1342da48ffb0 |
| ENCFF765PON.tabix.bed.gz.dvc | 106.0 B | 79cf8726b3bd6742c018ed3aad442c3d |
| ENCFF765PON.tabix.bed.gz.tbi | 135.63 KB | 3fd27d0bc2d867ca90025268c94f0d66 |
| ENCFF765PON.tabix.bed.gz.tbi.dvc | 110.0 B | 13130ef0bab2e00efd97e63f9363ceea |
| genomic_resource.yaml | 2.98 KB | 33e7d287eacf637d243fc0763ba72655 |
| genomic_resource_original.yaml | 2.88 KB | 9b2e4f925cbee4ab329de991dfe2e40b |
| statistics/ |