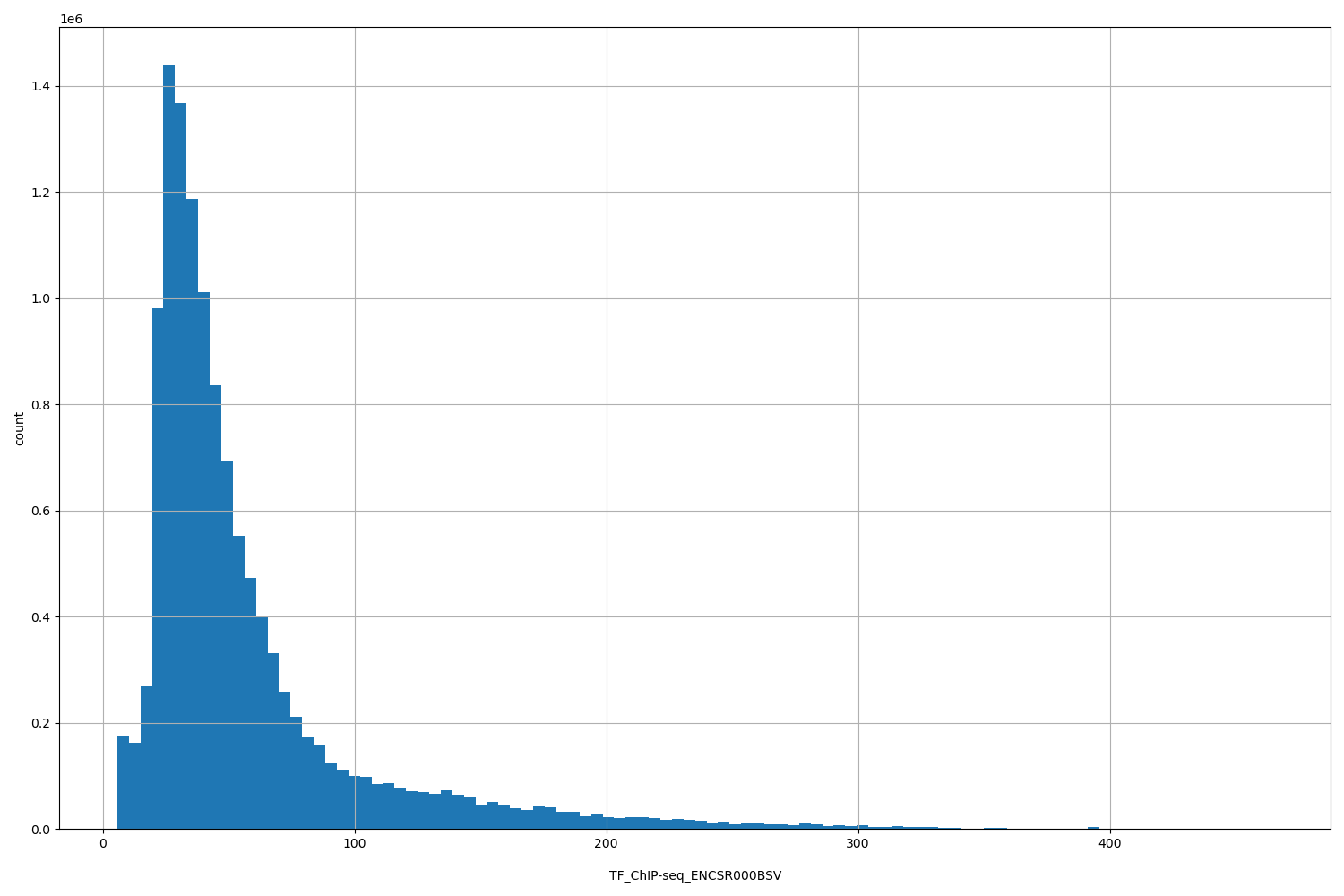
TF_ChIP-seq_ENCSR000BSV
| Id: | TF_ChIP-seq/ENCSR000BSV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BSV [biosamplesummary="Homo sapiens SK-N-SH" and target="NFIC"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN272YMO|/analyses/ENCAN272YMO/} has in progress subobject document {ec9c2509-773c-4002-91ab-4b84ceaf3483|/documents/ec9c2509-773c-4002-91ab-4b84ceaf3483/} audit_internal_action: Released analysis {ENCAN272YMO|/analyses/ENCAN272YMO/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF749DWZ|/files/ENCFF749DWZ/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.11. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF965AKM|/files/ENCFF965AKM/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.11. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BSV | float |
TF_ChIP-seq_ENCSR000BSV |
TF_ChIP-seq ENCSR000BSV [biosample_summary="Homo sapiens SK-N-SH" and target="NFIC"]
|
 |
[5.82, 465] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF749DWZ.bed.gz | 677.89 KB | 207edb94d5b6ebad31806f6105766a7d |
| ENCFF749DWZ.bed.gz.dvc | 100.0 B | 3da2881fcc3602fd6e423b764b0f3196 |
| ENCFF749DWZ.tabix.bed.gz | 490.03 KB | 3f9dbcdfed2c9537fc17ea745c4e61e3 |
| ENCFF749DWZ.tabix.bed.gz.dvc | 106.0 B | 7f159680a6d03f705145be0b760625b3 |
| ENCFF749DWZ.tabix.bed.gz.tbi | 219.31 KB | c1ebef244904cf89afec137e17904565 |
| ENCFF749DWZ.tabix.bed.gz.tbi.dvc | 110.0 B | f08fd4d7aebcca5f69ad71e32f985e19 |
| genomic_resource.yaml | 2.64 KB | 4eb4b01d8cf99fcc6639f2f6ba7ac5b6 |
| genomic_resource_original.yaml | 2.54 KB | 43d808bae055a4c25aac2df215daca5d |
| statistics/ |