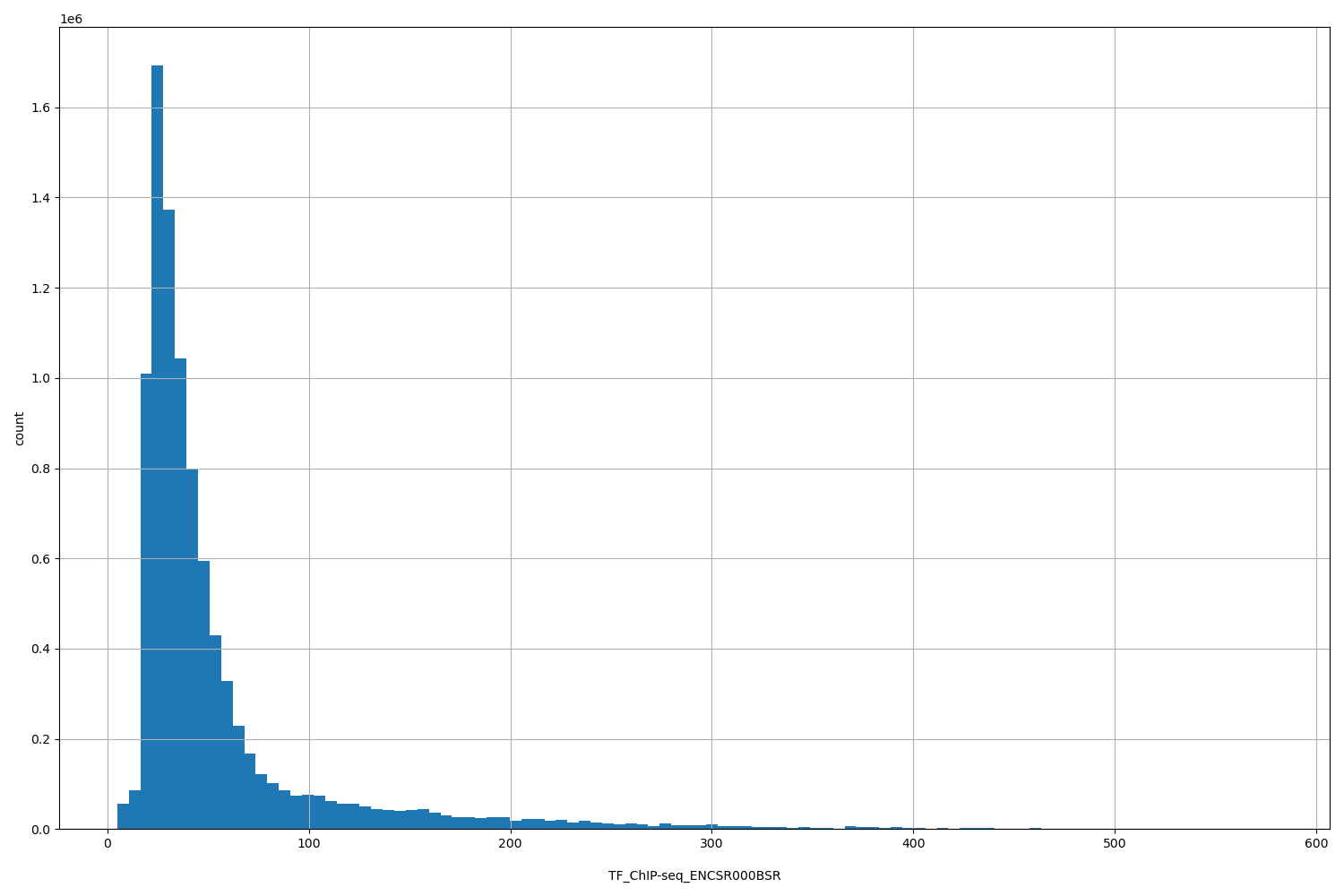
TF_ChIP-seq_ENCSR000BSR
| Id: | TF_ChIP-seq/ENCSR000BSR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BSR [biosamplesummary="Homo sapiens MCF-7" and target="CEBPB"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN087WJW|/analyses/ENCAN087WJW/} has in progress subobject document {c7373d0f-ddcc-406a-83d9-3e034fbd442f|/documents/c7373d0f-ddcc-406a-83d9-3e034fbd442f/} audit_internal_action: Released analysis {ENCAN087WJW|/analyses/ENCAN087WJW/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF341SFE|/files/ENCFF341SFE/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13780448 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting CEBPB-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF240ZHW|/files/ENCFF240ZHW/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.43 and a self consistency ratio of 2.27. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BSR | float |
TF_ChIP-seq_ENCSR000BSR |
TF_ChIP-seq ENCSR000BSR [biosample_summary="Homo sapiens MCF-7" and target="CEBPB"]
|
 |
[4.84, 578] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF240ZHW.bed.gz | 641.57 KB | 8c20c751b18b07075d20c6b9339c3dd4 |
| ENCFF240ZHW.bed.gz.dvc | 100.0 B | 624078f21da73f83f21271740a3a06c1 |
| ENCFF240ZHW.tabix.bed.gz | 442.3 KB | 7b2fcc0192ad2d96c46554ff0d6864ea |
| ENCFF240ZHW.tabix.bed.gz.dvc | 106.0 B | e6e8230a40c9979801892ce9cb1cf065 |
| ENCFF240ZHW.tabix.bed.gz.tbi | 238.91 KB | efb042aeb50168b0245248b210dda2df |
| ENCFF240ZHW.tabix.bed.gz.tbi.dvc | 110.0 B | db8a8cf5d5262fa4776026f186c0640d |
| genomic_resource.yaml | 2.6 KB | edecd0bb619d97d9b10dfe3b38b542c7 |
| genomic_resource_original.yaml | 2.51 KB | 409a877225ac355f1e5162335c4cc386 |
| statistics/ |