TF_ChIP-seq_ENCSR000BRZ
| Id: | TF_ChIP-seq/ENCSR000BRZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BRZ [biosamplesummary="Homo sapiens HCT116" and target="EGR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN541DZY|/analyses/ENCAN541DZY/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF329JMS|/files/ENCFF329JMS/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 10280579 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EGR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
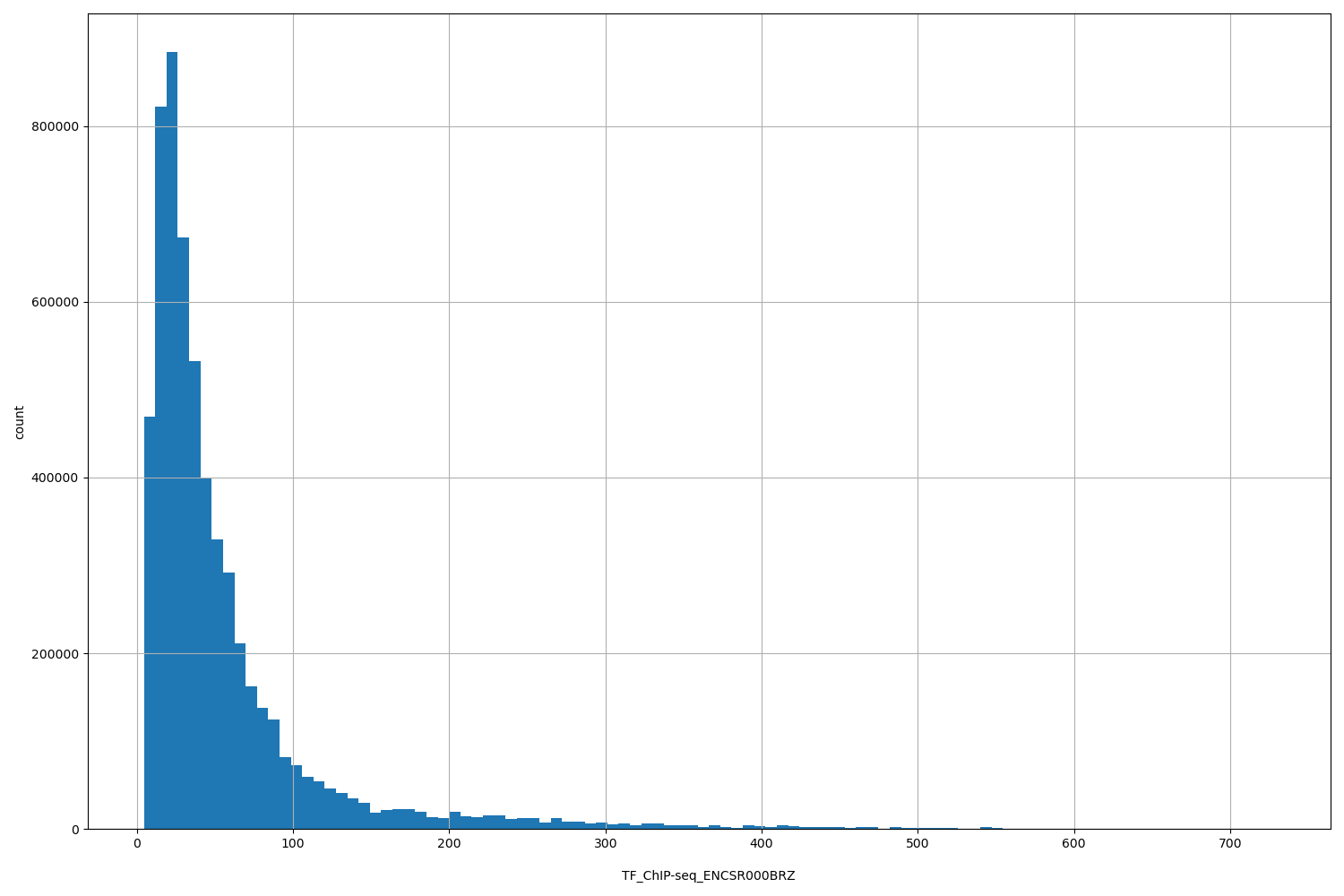
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BRZ | float |
TF_ChIP-seq_ENCSR000BRZ |
TF_ChIP-seq ENCSR000BRZ [biosample_summary="Homo sapiens HCT116" and target="EGR1"]
|
 |
[4.48, 728] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF822VKJ.bed.gz | 263.97 KB | 4b9821a3e4f6b7ff9786a9a9558dc7e7 |
| ENCFF822VKJ.bed.gz.dvc | 100.0 B | 9dccee8e4bf2e94e79fc2eeb7981c39e |
| ENCFF822VKJ.tabix.bed.gz | 201.94 KB | e055800d8fc912553525458957ad5a20 |
| ENCFF822VKJ.tabix.bed.gz.dvc | 106.0 B | 4ee22ab93cdb2b90c120b23139dbf858 |
| ENCFF822VKJ.tabix.bed.gz.tbi | 102.04 KB | 1cf5859664fedcb4cc2c517d7ff132ab |
| ENCFF822VKJ.tabix.bed.gz.tbi.dvc | 110.0 B | 07d391cd53952d01c7961b0ebe12a838 |
| genomic_resource.yaml | 1.84 KB | b89f514d255c6b6d9f67b31e2263ca3e |
| genomic_resource_original.yaml | 1.74 KB | 854167895ae85932e1feac173bb98607 |
| statistics/ |