TF_ChIP-seq_ENCSR000BRP
TF_ChIP-seq ENCSR000BRP [biosample_summary="Homo sapiens HepG2" and target="TEAD4"]
| Id: | TF_ChIP-seq/ENCSR000BRP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BRP [biosamplesummary="Homo sapiens HepG2" and target="TEAD4"] |
| Description: |
status: archived biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CLG|/files/ENCFF002CLG/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
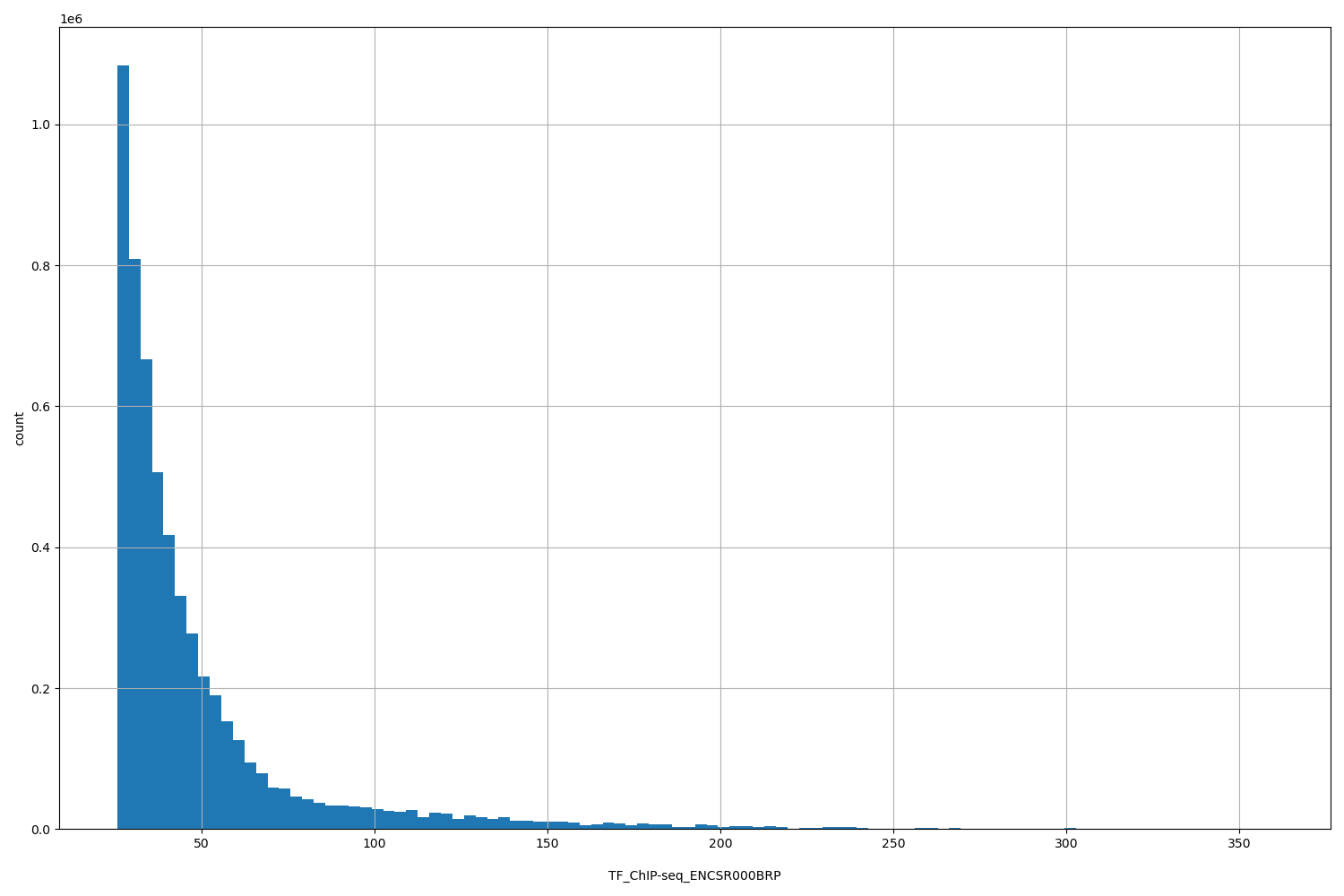
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BRP | float |
TF_ChIP-seq_ENCSR000BRP |
TF_ChIP-seq ENCSR000BRP [biosample_summary="Homo sapiens HepG2" and target="TEAD4"]
|
 |
[25.6, 360] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CLG.bed.gz | 291.74 KB | 4fedf810164ae7df7dd049dca4a257aa |
| ENCFF002CLG.bed.gz.dvc | 100.0 B | 4af54e30a805afd7e57d5d1877710d12 |
| ENCFF002CLG.tabix.bed.gz | 237.5 KB | 280fd8700d2e52757f9d22117d044d53 |
| ENCFF002CLG.tabix.bed.gz.dvc | 106.0 B | d9376d7016ae895d8cf0e8f3dd1b6294 |
| ENCFF002CLG.tabix.bed.gz.tbi | 109.4 KB | f989eddce8b4162f3385a5737c527f60 |
| ENCFF002CLG.tabix.bed.gz.tbi.dvc | 110.0 B | 9d7b3b7c93666dfee6b58d18f69561d9 |
| genomic_resource.yaml | 1.47 KB | 3880466161aaf831794163f4b673f147 |
| genomic_resource_original.yaml | 1.38 KB | abc14875a6bc772690fd03b529c7e2ef |
| statistics/ |