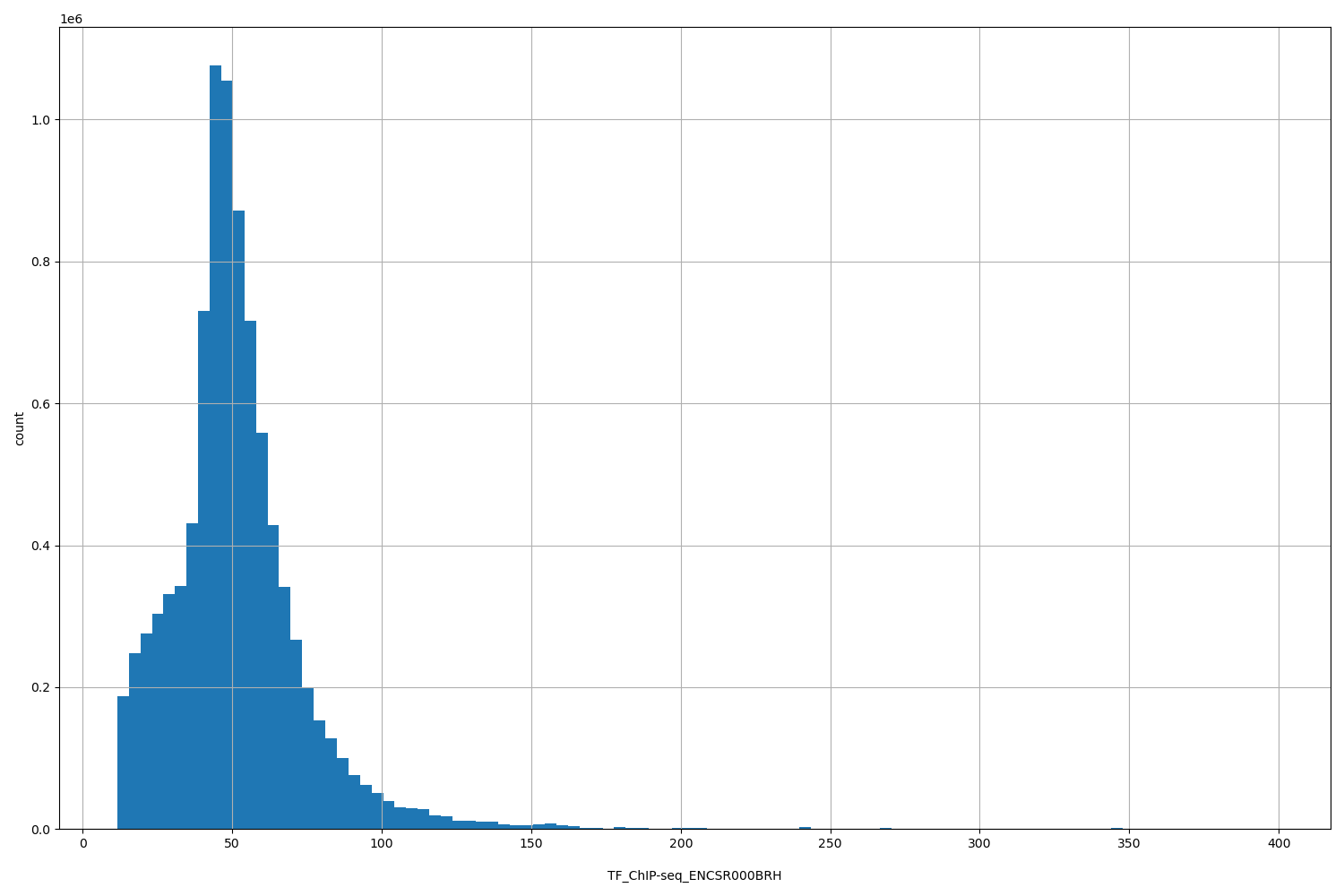
TF_ChIP-seq_ENCSR000BRH
| Id: | TF_ChIP-seq/ENCSR000BRH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BRH [biosamplesummary="Homo sapiens GM12878" and target="MTA3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN992QDM|/analyses/ENCAN992QDM/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF202XQV|/files/ENCFF202XQV/}, {ENCFF457WBH|/files/ENCFF457WBH/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.33 and a self consistency ratio of 3.57. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF413VUC|/files/ENCFF413VUC/}, {ENCFF611WSJ|/files/ENCFF611WSJ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.33 and a self consistency ratio of 3.57. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BRH | float |
TF_ChIP-seq_ENCSR000BRH |
TF_ChIP-seq ENCSR000BRH [biosample_summary="Homo sapiens GM12878" and target="MTA3"]
|
 |
[11.5, 398] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF413VUC.bed.gz | 315.37 KB | 986211e162ebe80002d15d053f4e37f8 |
| ENCFF413VUC.bed.gz.dvc | 100.0 B | 4d8ef807abb41c1dcb63a4389be29c91 |
| ENCFF413VUC.tabix.bed.gz | 250.58 KB | fcaaa41063ca97e8fdf2368b60a8e6a3 |
| ENCFF413VUC.tabix.bed.gz.dvc | 106.0 B | 3ecf65009d1fb3cbd57fbde96ef949f3 |
| ENCFF413VUC.tabix.bed.gz.tbi | 102.18 KB | 80dd41ff9cd5807d9306647f60129622 |
| ENCFF413VUC.tabix.bed.gz.tbi.dvc | 110.0 B | dbe01eeca28768e127faa8582783aa85 |
| genomic_resource.yaml | 2.48 KB | 3cca2808f08c5dada1a7c3c5b7d2fa4c |
| genomic_resource_original.yaml | 2.38 KB | 8accfcdb6fbd4fd4cba293377d476a67 |
| statistics/ |