TF_ChIP-seq_ENCSR000BKM
TF_ChIP-seq ENCSR000BKM [biosample_summary="Homo sapiens K562" and target="GATA2"]
| Id: | TF_ChIP-seq/ENCSR000BKM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BKM [biosamplesummary="Homo sapiens K562" and target="GATA2"] |
| Description: |
status: archived biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CMA|/files/ENCFF002CMA/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
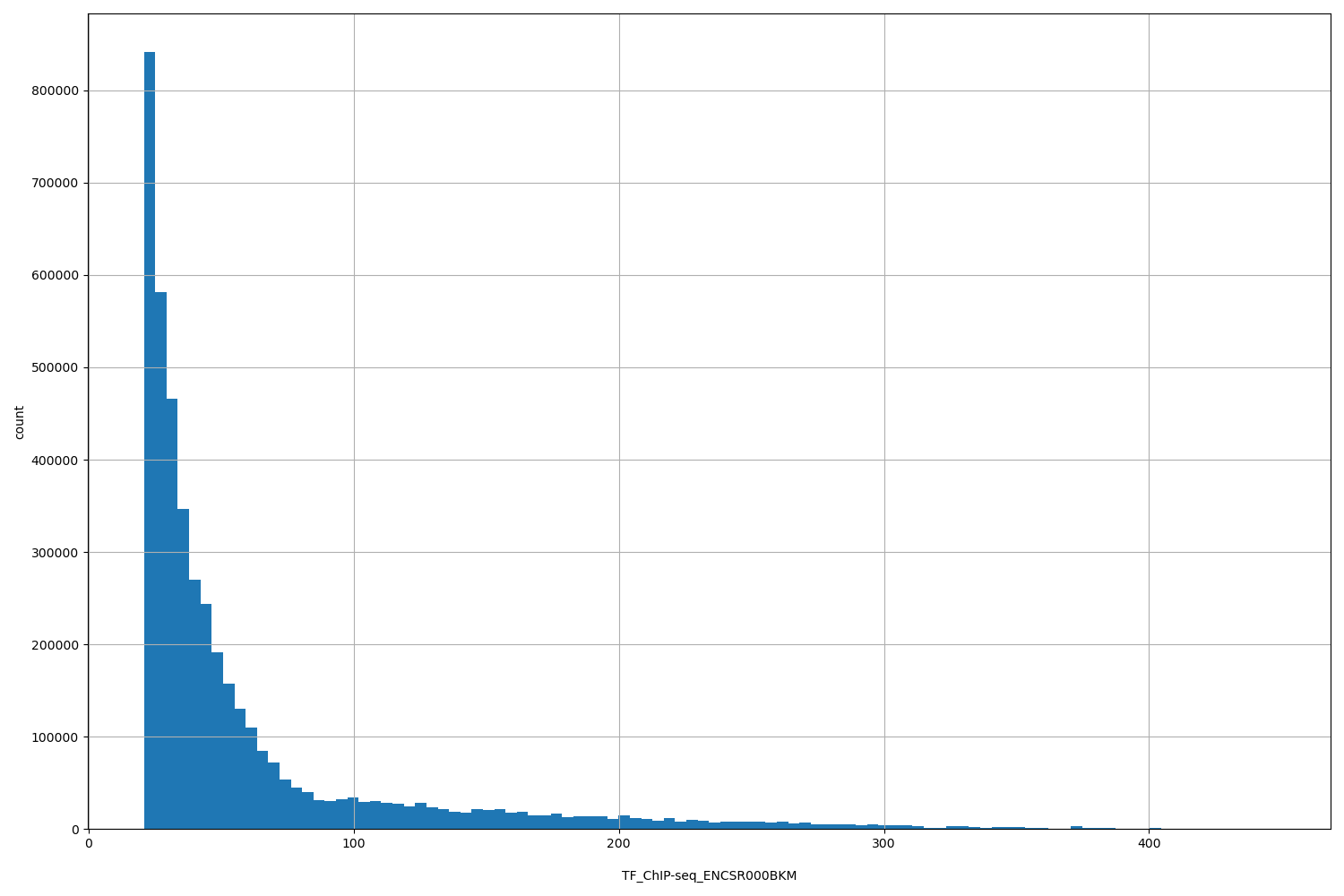
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BKM | float |
TF_ChIP-seq_ENCSR000BKM |
TF_ChIP-seq ENCSR000BKM [biosample_summary="Homo sapiens K562" and target="GATA2"]
|
 |
[20.9, 447] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CMA.bed.gz | 370.44 KB | 375a3d9a300928e6b93f233463063ae9 |
| ENCFF002CMA.bed.gz.dvc | 100.0 B | 27c0d892c9820ffb11ef21da9b47bff8 |
| ENCFF002CMA.tabix.bed.gz | 294.04 KB | 0a38285f963c7c4a149d15430c927261 |
| ENCFF002CMA.tabix.bed.gz.dvc | 106.0 B | ece3ddf6ac2fbfecdbc62862bc28859e |
| ENCFF002CMA.tabix.bed.gz.tbi | 132.85 KB | 7fddd2a974cc67ba81c5dde60f5be704 |
| ENCFF002CMA.tabix.bed.gz.tbi.dvc | 110.0 B | 921bc107b69ba215707f26c54f13bfa3 |
| genomic_resource.yaml | 1.47 KB | e1aabddd0caf158aac32934557452498 |
| genomic_resource_original.yaml | 1.38 KB | 09697a06ced0913933bfb0c2f4480475 |
| statistics/ |