TF_ChIP-seq_ENCSR000BJJ
| Id: | TF_ChIP-seq/ENCSR000BJJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BJJ [biosamplesummary="Homo sapiens SK-N-SH" and target="REST"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN120BZI|/analyses/ENCAN120BZI/} has in progress subobject document {35f4af5f-3e14-482f-bb41-dbd1fd1524a7|/documents/35f4af5f-3e14-482f-bb41-dbd1fd1524a7/} audit_internal_action: Released analysis {ENCAN120BZI|/analyses/ENCAN120BZI/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF358XFJ|/files/ENCFF358XFJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 3.68 and a self consistency ratio of 5.56. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Processed alignments file {ENCFF310DLW|/files/ENCFF310DLW/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 10566392 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting REST-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF155PZC|/files/ENCFF155PZC/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13651188 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting REST-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
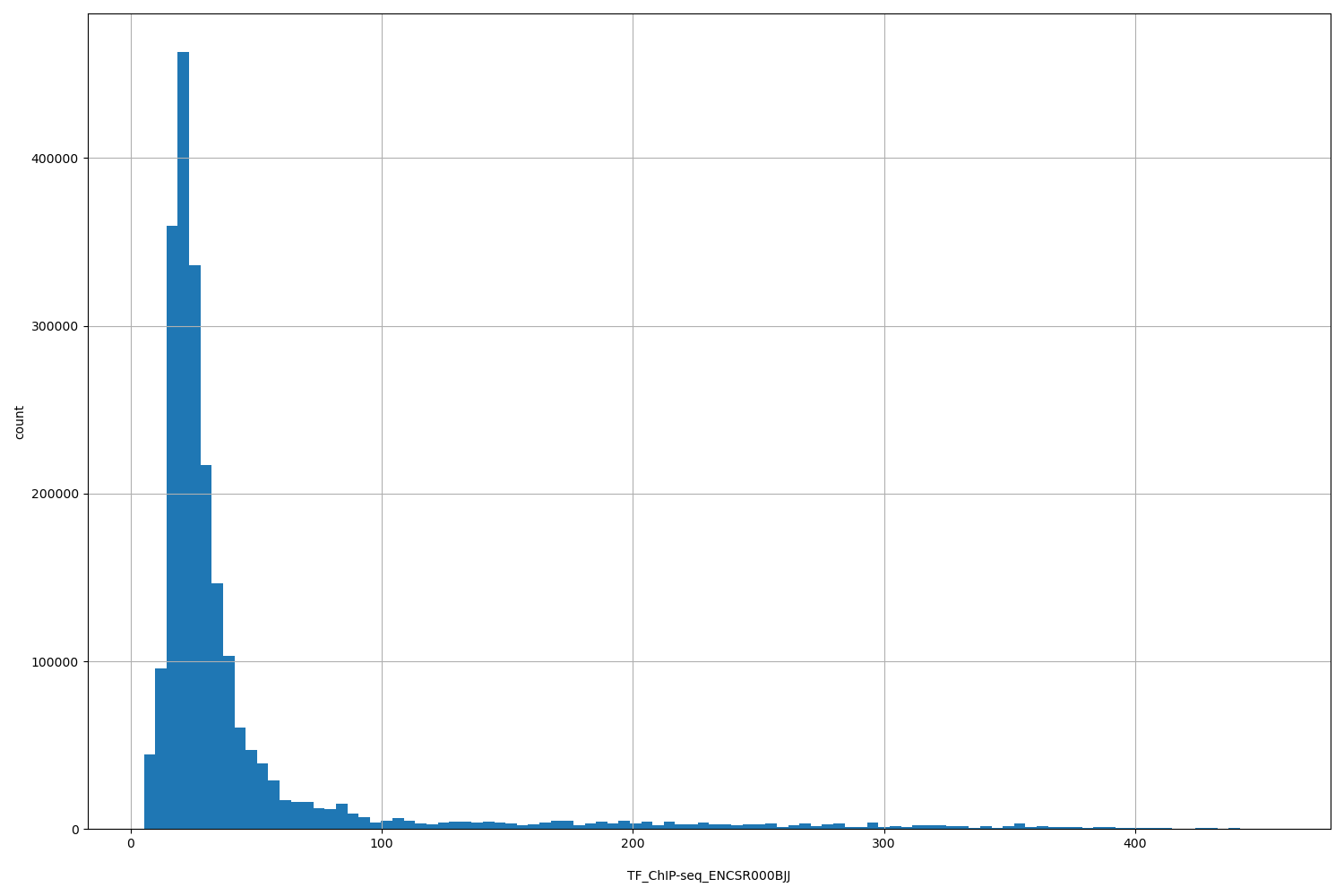
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BJJ | float |
TF_ChIP-seq_ENCSR000BJJ |
TF_ChIP-seq ENCSR000BJJ [biosample_summary="Homo sapiens SK-N-SH" and target="REST"]
|
 |
[5.25, 455] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF358XFJ.bed.gz | 172.2 KB | c05cae43de81338b910ee2ee56733ee4 |
| ENCFF358XFJ.bed.gz.dvc | 100.0 B | 0dc329564d9fdc3f55c827969645cde3 |
| ENCFF358XFJ.tabix.bed.gz | 113.22 KB | d5374484c85b7ef382ca47a5bc998a27 |
| ENCFF358XFJ.tabix.bed.gz.dvc | 106.0 B | 8e203f82fef22ba990ff58871f2dc16d |
| ENCFF358XFJ.tabix.bed.gz.tbi | 79.78 KB | 3a6224e658ae6d5e6784b935f9edb351 |
| ENCFF358XFJ.tabix.bed.gz.tbi.dvc | 109.0 B | b63236dd0e68cfe0c4b419835cba129a |
| genomic_resource.yaml | 3.26 KB | ad41347801b19c4bb993f317feafcfbc |
| genomic_resource_original.yaml | 3.16 KB | f2a940204b63a1b03b6908a734c0cf13 |
| statistics/ |