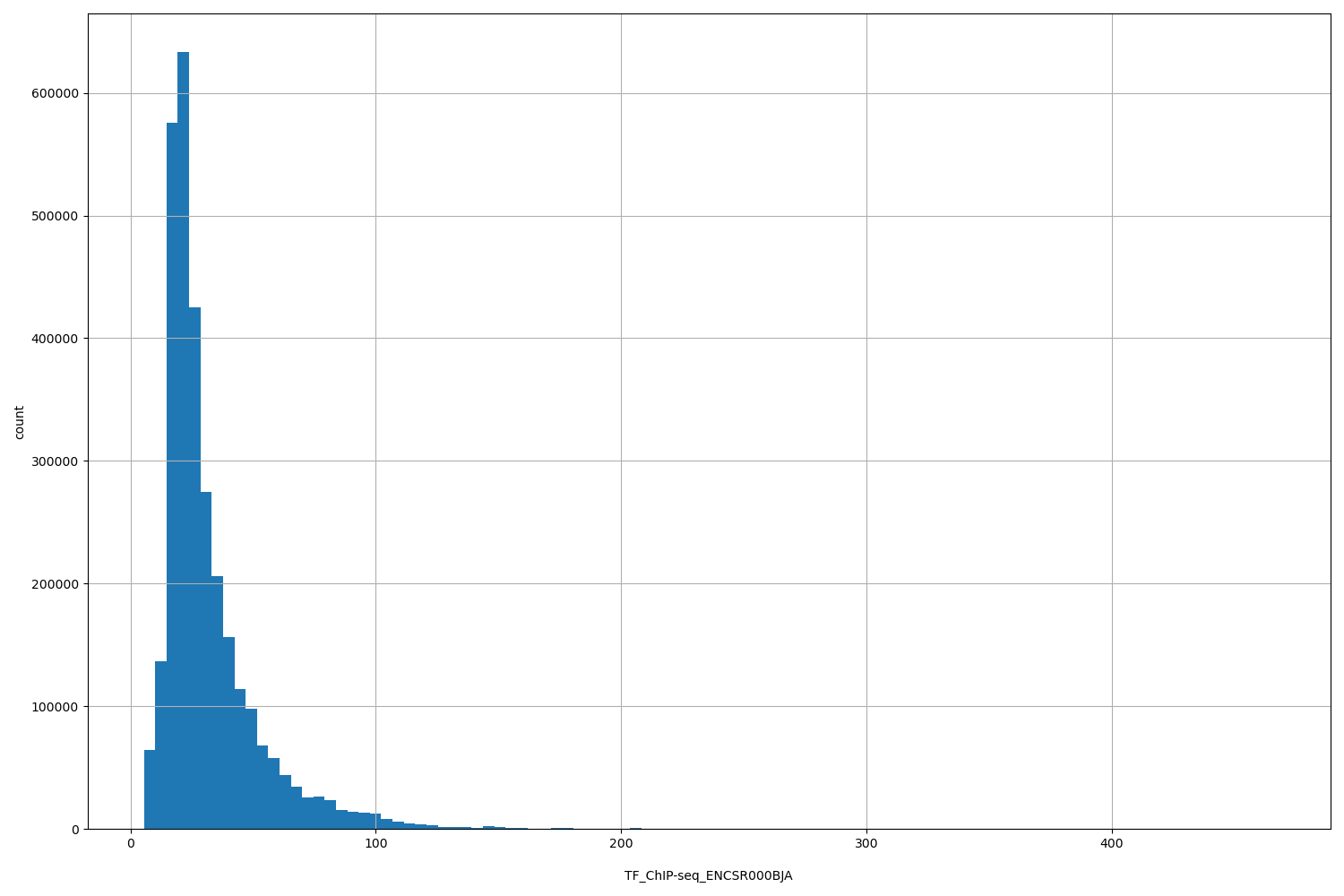
TF_ChIP-seq_ENCSR000BJA
| Id: | TF_ChIP-seq/ENCSR000BJA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BJA [biosamplesummary="Homo sapiens H1" and target="EGR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF477ANT|/files/ENCFF477ANT/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN287KFQ|/analyses/ENCAN287KFQ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF777MZR|/files/ENCFF777MZR/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 17905180 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EGR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF054DVF|/files/ENCFF054DVF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 19044316 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting EGR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF004MNA|/files/ENCFF004MNA/}, {ENCFF477ANT|/files/ENCFF477ANT/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.46. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF469MQG|/files/ENCFF469MQG/}, {ENCFF042WRS|/files/ENCFF042WRS/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.28 and a self consistency ratio of 2.46. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BJA | float |
TF_ChIP-seq_ENCSR000BJA |
TF_ChIP-seq ENCSR000BJA [biosample_summary="Homo sapiens H1" and target="EGR1"]
|
 |
[5.47, 466] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF477ANT.bed.gz | 200.72 KB | 29679d804ec39d878f61ec037a69b2b7 |
| ENCFF477ANT.bed.gz.dvc | 100.0 B | 2aa4ea874e65e3133eef85792af890b8 |
| ENCFF477ANT.tabix.bed.gz | 140.57 KB | 56fc851d41a16b46a67aa35fe1169b3a |
| ENCFF477ANT.tabix.bed.gz.dvc | 106.0 B | 4658c8934f8f65c7a7c5a9b82b17a832 |
| ENCFF477ANT.tabix.bed.gz.tbi | 91.7 KB | 1bc7d36dfde3d9997ee96d0cf9e6a7e7 |
| ENCFF477ANT.tabix.bed.gz.tbi.dvc | 109.0 B | 873c8ad713ef2de4ae0e233c75897de7 |
| genomic_resource.yaml | 3.64 KB | cc179a318a0320d3d5aa5588a0ccc562 |
| genomic_resource_original.yaml | 3.55 KB | 23ee0966e3699e2a028688e3bef50e00 |
| statistics/ |