TF_ChIP-seq_ENCSR000BIJ
| Id: | TF_ChIP-seq/ENCSR000BIJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BIJ [biosamplesummary="Homo sapiens GM12891" and target="SPI1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN442JLB|/analyses/ENCAN442JLB/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF744AGB|/files/ENCFF744AGB/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: Processed alignments file {ENCFF482TVZ|/files/ENCFF482TVZ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 13427665 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SPI1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF724WMD|/files/ENCFF724WMD/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 16501012 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SPI1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
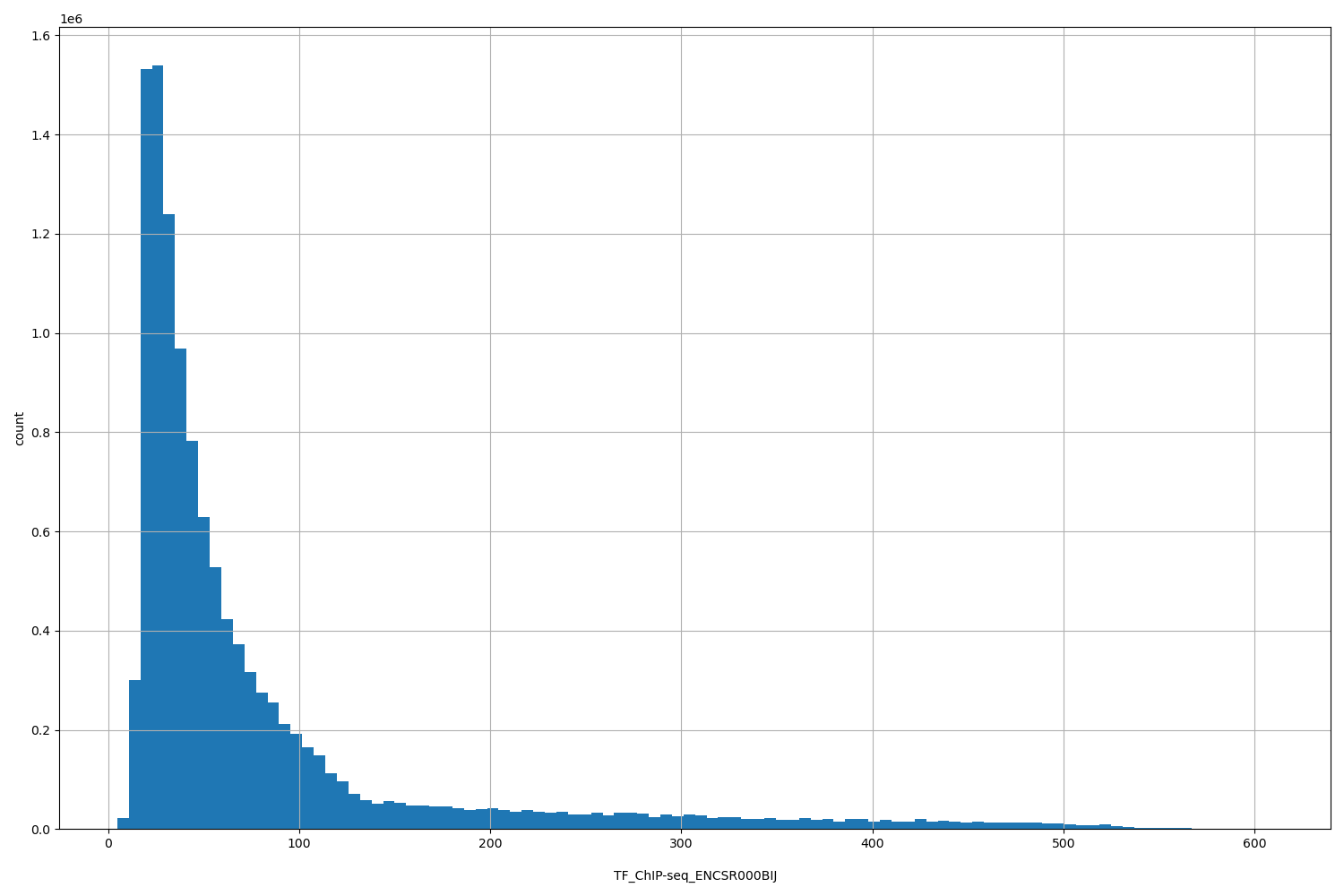
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BIJ | float |
TF_ChIP-seq_ENCSR000BIJ |
TF_ChIP-seq ENCSR000BIJ [biosample_summary="Homo sapiens GM12891" and target="SPI1"]
|
 |
[4.76, 610] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF744AGB.bed.gz | 902.4 KB | 8bb508bd4bb040c539b1c331dd860997 |
| ENCFF744AGB.bed.gz.dvc | 100.0 B | f42d7bea2e8192e9fdda0ea44c3a1671 |
| ENCFF744AGB.tabix.bed.gz | 621.43 KB | 166e20db5a44626f757b10e9e6bf0da1 |
| ENCFF744AGB.tabix.bed.gz.dvc | 106.0 B | 39632b6939eb790f90777cadc420f958 |
| ENCFF744AGB.tabix.bed.gz.tbi | 295.54 KB | 3561962f3bab930c2ca9a24d21932156 |
| ENCFF744AGB.tabix.bed.gz.tbi.dvc | 110.0 B | 0f1fe8f04351486c5dc054137083dfff |
| genomic_resource.yaml | 2.52 KB | 0861ebb6990662e83ddc3903e9d8131a |
| genomic_resource_original.yaml | 2.43 KB | ec0cc171a2780699430ff4afeacc0bf7 |
| statistics/ |