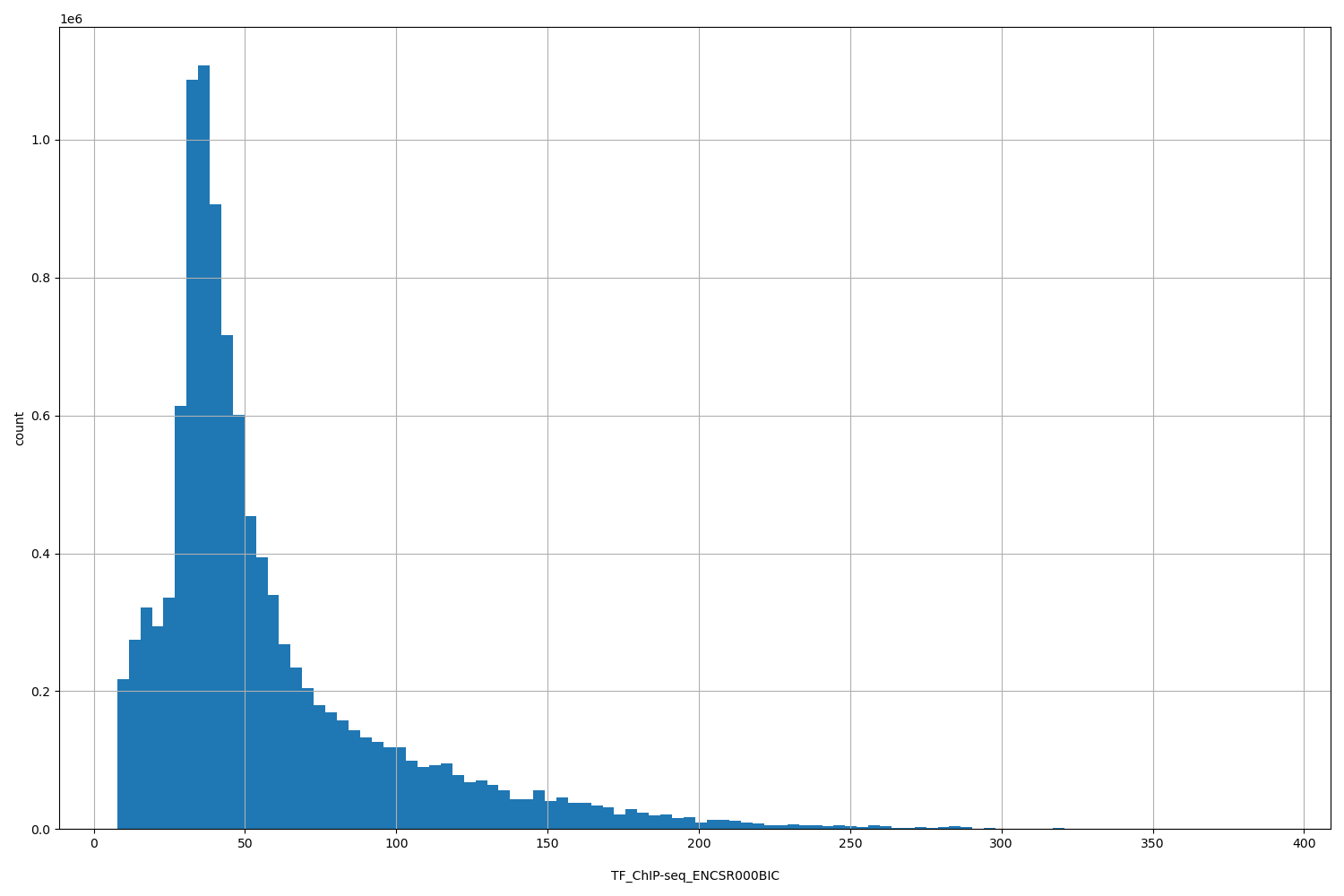
TF_ChIP-seq_ENCSR000BIC
| Id: | TF_ChIP-seq/ENCSR000BIC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BIC [biosamplesummary="Homo sapiens H1" and target="POLR2AphosphoS5"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF418QVJ|/files/ENCFF418QVJ/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN228YRJ|/analyses/ENCAN228YRJ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF079DUB|/files/ENCFF079DUB/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 14030294 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2AphosphoS5-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BIC | float |
TF_ChIP-seq_ENCSR000BIC |
TF_ChIP-seq ENCSR000BIC [biosample_summary="Homo sapiens H1" and target="POLR2AphosphoS5"]
|
 |
[7.78, 390] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF418QVJ.bed.gz | 570.8 KB | e02c29a59b2bea50cecbdaf1d09060a2 |
| ENCFF418QVJ.bed.gz.dvc | 100.0 B | 215ca4b8f8f24b4848d0c60e0368913c |
| ENCFF418QVJ.tabix.bed.gz | 427.12 KB | 30a363b5b2928020f0dd6d83069ebcbe |
| ENCFF418QVJ.tabix.bed.gz.dvc | 106.0 B | bb216978a3777980b8739c5063047bf7 |
| ENCFF418QVJ.tabix.bed.gz.tbi | 141.88 KB | 68a30e9c2d8e614d926c43c21a21d450 |
| ENCFF418QVJ.tabix.bed.gz.tbi.dvc | 110.0 B | fed81c912ef14dec3d8b9d5ea8d36f64 |
| genomic_resource.yaml | 2.06 KB | 1b7d6c81ae821692c3b155d70a42c385 |
| genomic_resource_original.yaml | 1.96 KB | eac8332f5deb3ddb9ad565c926e1b670 |
| statistics/ |