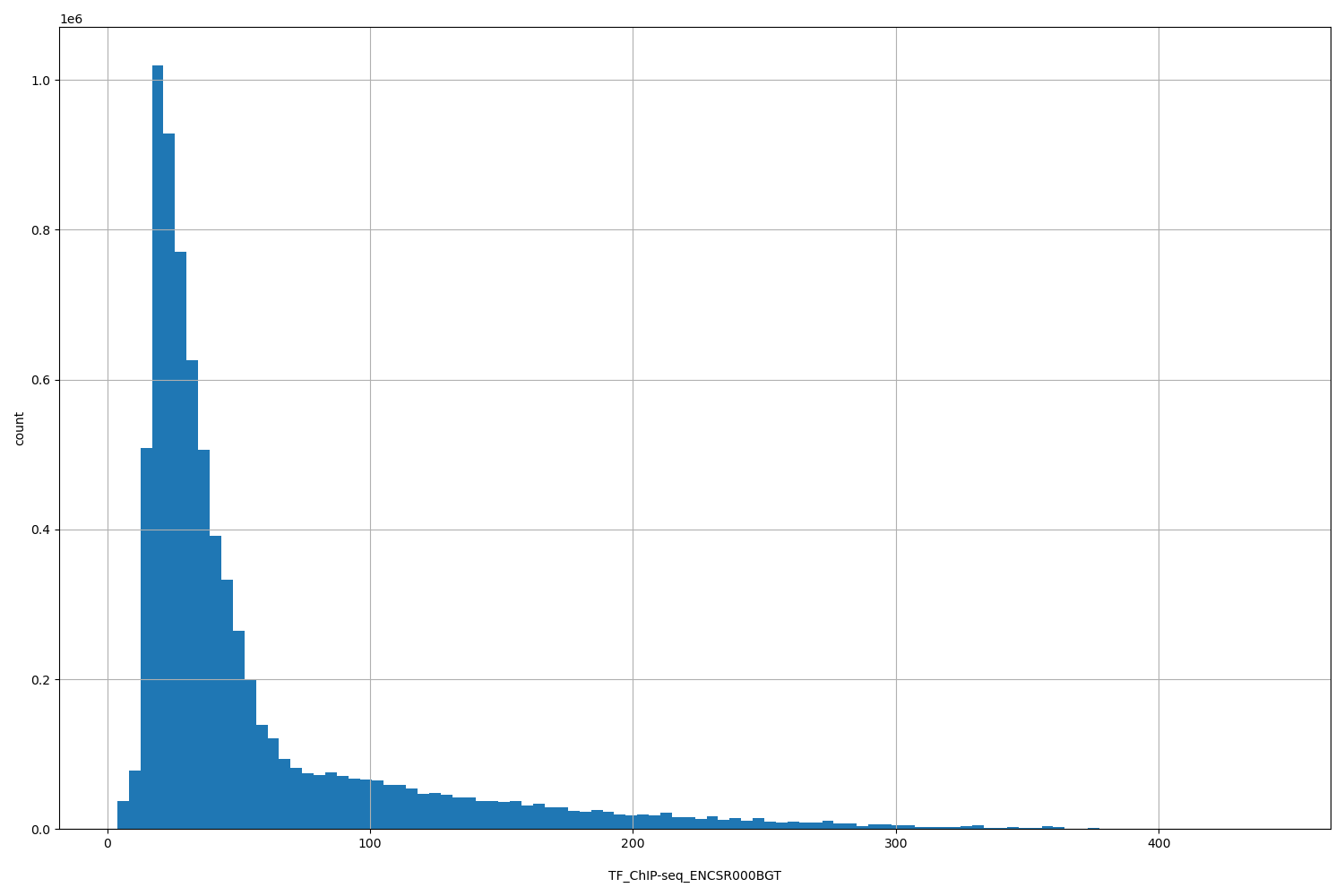
TF_ChIP-seq_ENCSR000BGT
| Id: | TF_ChIP-seq/ENCSR000BGT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000BGT [biosamplesummary="Homo sapiens GM12878" and target="BATF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN915BOG|/analyses/ENCAN915BOG/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN915BOG|/analyses/ENCAN915BOG/} has in progress subobject document {3b9ad243-12fd-4210-842b-ecd55ff06efa|/documents/3b9ad243-12fd-4210-842b-ecd55ff06efa/} audit_warning: Processed alignments file {ENCFF859PRS|/files/ENCFF859PRS/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 13949632 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BATF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF755RCM|/files/ENCFF755RCM/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 15104537 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting BATF-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000BGT | float |
TF_ChIP-seq_ENCSR000BGT |
TF_ChIP-seq ENCSR000BGT [biosample_summary="Homo sapiens GM12878" and target="BATF"]
|
 |
[3.83, 443] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF728KFD.bed.gz | 569.45 KB | 7f22a3244fa244a435a39df67ec6a4f0 |
| ENCFF728KFD.bed.gz.dvc | 100.0 B | 95e06b7b444a4645e441aa86760293ef |
| ENCFF728KFD.tabix.bed.gz | 431.18 KB | 65b004d3f8b9f1f970b7f4929b861f07 |
| ENCFF728KFD.tabix.bed.gz.dvc | 106.0 B | 778440c62bc898d0b1f2a03f1a31c77d |
| ENCFF728KFD.tabix.bed.gz.tbi | 219.54 KB | 55a2c0c78d076db3a2765c8366c8f5cf |
| ENCFF728KFD.tabix.bed.gz.tbi.dvc | 110.0 B | 8fd16292c2e94a9d2f234ad297807b83 |
| genomic_resource.yaml | 2.56 KB | 1972993c87a145fc3a92366ba47a0321 |
| genomic_resource_original.yaml | 2.47 KB | 59d6776b461c28538d406d36ce5a6f28 |
| statistics/ |